Swiss Year of Scientometrics Lecture: Opportunities and Challenges of Scientometrics–Part IV
This blog post is the last of a four part series based on the keynote presentation by Stefanie Haustein at the Swiss Year of Scientometrics lecture and workshop series at ETH Zurich on June 7, 2023.

Responsible use of metrics
As a last topic, I want to address a very important area that I think for a very long time has been ignored by the scientometric community—namely the use of the metrics that it creates and computes.
Research assessment reform
We all know that we live in a “publish or perish” culture, much of which has been created by quantitative measurements of scientific productivity and impact.
Metrics—such as the journal's impact factor (JIF) and h-index—are used to make critical decisions about individuals, including review, promotion, tenure (RPT), funding, and academic awards. The importance of quantitative metrics has changed scholarly communication and publishing, and created a range of adverse effects, such as plagiarism, “salami publishing,” authorship for sale, and citation cartels.
These adverse effects can be described by Goodhart’s and Campbell’s laws. By setting a metric as a target and by linking that target to a reward structure, we create an incentive for that metric to be gamed.
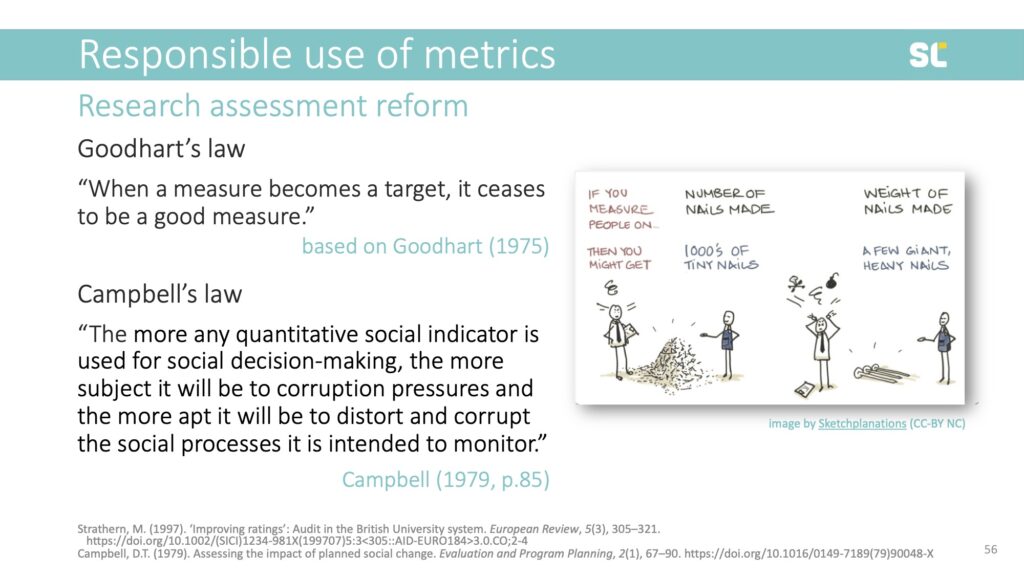

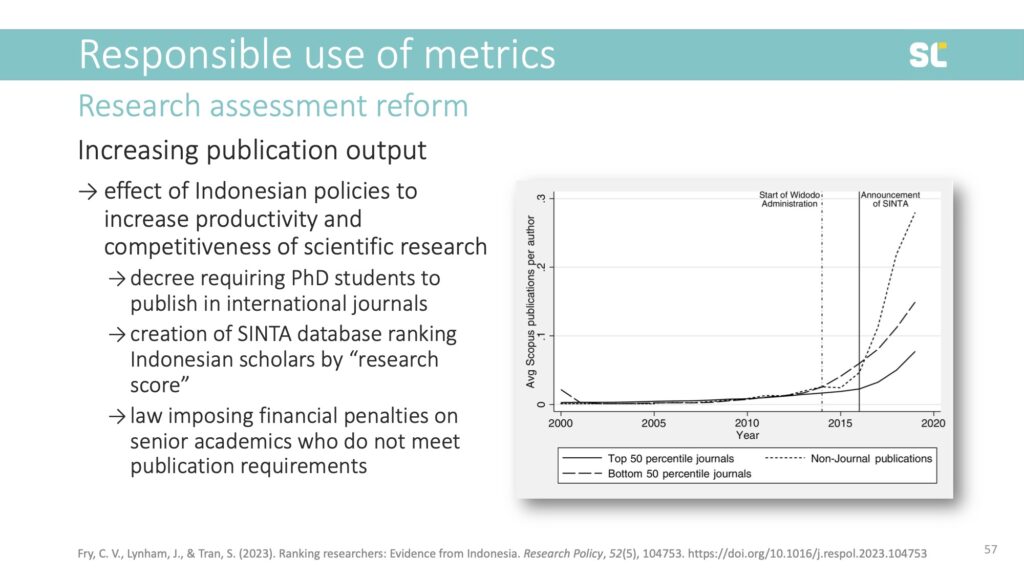
There are some examples of quantitative metrics that have actually reached desired goals of improving research productivity. A recent example comes from Indonesia, where policies instated by the Widodo administration increased the number of publications in Scopus-indexed journals.
The policies included a decree that requires PhD students to publish one paper in an international journal, the creation of the SINTA database indexing Indonesian publications and ranking scholars by a “research score,” and even a law imposing financial penalties on senior academics who do not meet a certain publication output. Below, you clearly see the increase in publication output after the policy change.
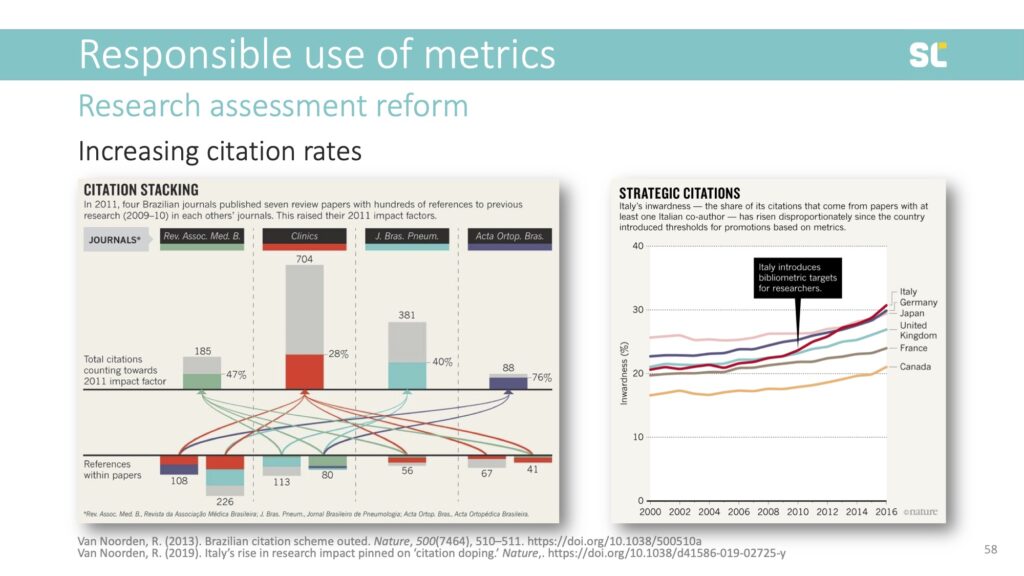
Unfortunately, we frequently see issues where researchers try to game metrics and metrics cause so-called adverse effects. In the slide below, you see an example of “citation stacking” (left image)—where journals try to increase their impact factors by citing each other’s papers published in the two previous years. Citation databases now monitor stacking and suspend journals. Another example is “citation doping” (right image), where you could see a significant rise in self-citations after Italy introduced bibliometric targets for researchers in 2010.
Since the current form of research assessment is harming research and researchers in all disciplines, there are some initiatives that are trying to reform it:
- The San Francisco Declaration on Research Assessment (DORA) is now turning 10 years
- The Leiden Manifesto for Research Metrics was launched in 2015 by the scientometric community
- The Coalition for Advancing Research Assessment (CoARA) was formed in 2022
For example, CoARA’s core commitments include using quantitative indicators only as a supplement to support peer review, abandoning inappropriate use of metrics (particularly the JIF and h-index), and avoiding the use of university rankings.
CoARA also prominently stresses the importance of evaluating practices, criteria, and tools of research assessment using solid evidence and state-of-the-art research on research, and making data openly available for evidence gathering and research.
These are all steps in the right direction but behavior change can take a long time, particularly in academia.
Metrics Literacies
Lastly, I want to talk about how we can educate researchers and research administrators about responsible use of scientometric indicators.
I want to introduce what we call “Metrics Literacies,” an integrated set of competencies, dispositions, and knowledge that empower individuals to recognize, interpret, critically assess, and effectively and ethically use scholarly metrics.
While Metrics Literacies are essential, there is a lack of effective and efficient educational resources. We argue that we need to educate researchers and research administrators about scholarly metrics.
In the context where every researcher is already overwhelmed with keeping up with the literature in their own fields, videos are more efficient and effective than text. Storytelling and personas can help to draw attention and increase retention.

So we are currently planning a randomized controlled trial to test the effectiveness of different types of educational videos, using the h-index as an example.
Before producing our own educational videos about the h-index, we assessed the landscape of existing videos on Youtube, reviewing 274 videos and coding 31 that met the inclusion criteria according to a codebook with 50 variables.

We used Cognitive Load Theory to assess the videos:
- Extraneous cognitive load requires cognitive processing that is not related to learning and should be reduced. This includes non-productive distractions (e.g., background noise).
- Intrinsic cognitive load is associated with inherently difficult tasks or topics, this burden can be reduced by breaking down content (e.g., segmenting).
- Germane cognitive load refers to the resources required to facilitate learning, which can be enhanced by presenting content multi-modally (e.g., dual channel).
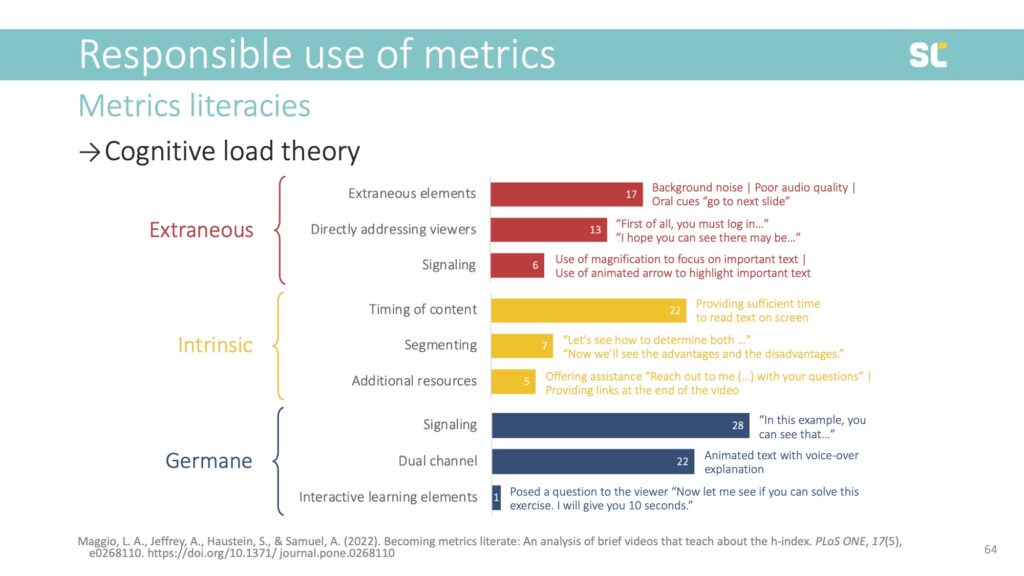
We found that among existing videos about the h-index, 22% do not define the metric and almost half do not address any of its caveats. For example, 61% do not address disciplinary differences and 77% do not mention its inability to assess early career researchers. This is highly problematic because it further promotes the inappropriate use of this flawed indicator.
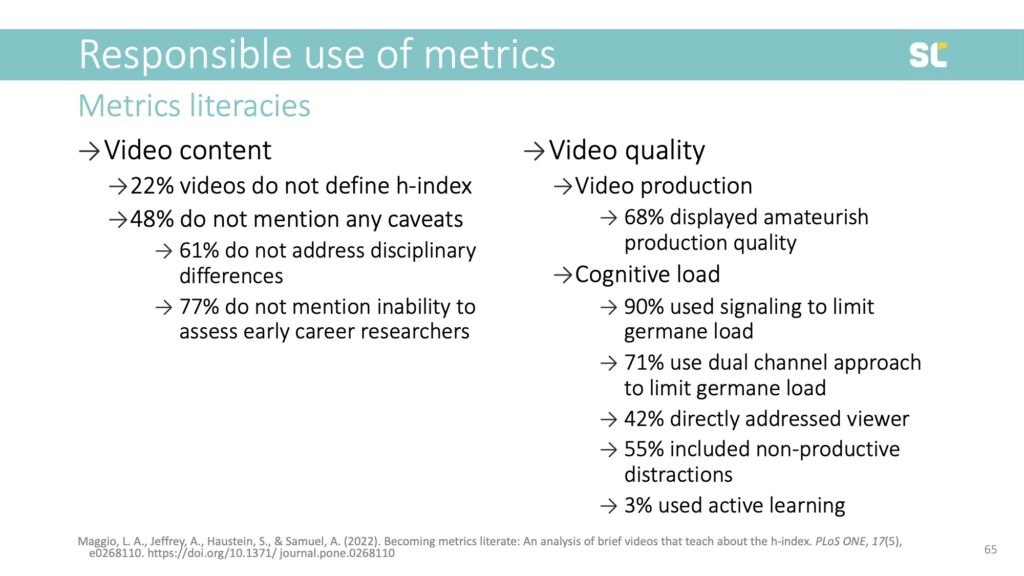

From the perspective of production quality, we found that most videos seemed amateurish and more than half contained non-productive distractions. Only one of the 31 coded videos used active learning elements.
We are currently preparing the production of three videos about the h-index, using different educational video formats (e.g., talking head and animations) to set up a randomized controlled trial to test their effectiveness against the control—a regular academic text, which represents the traditional modality of scholarly communication but is believed to be less efficient and engaging as an educational resource.
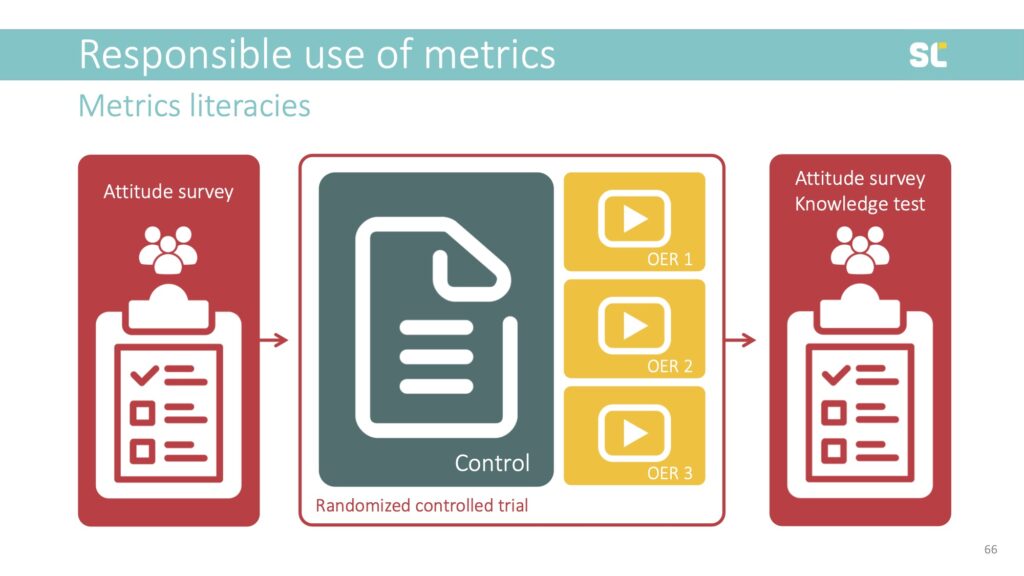
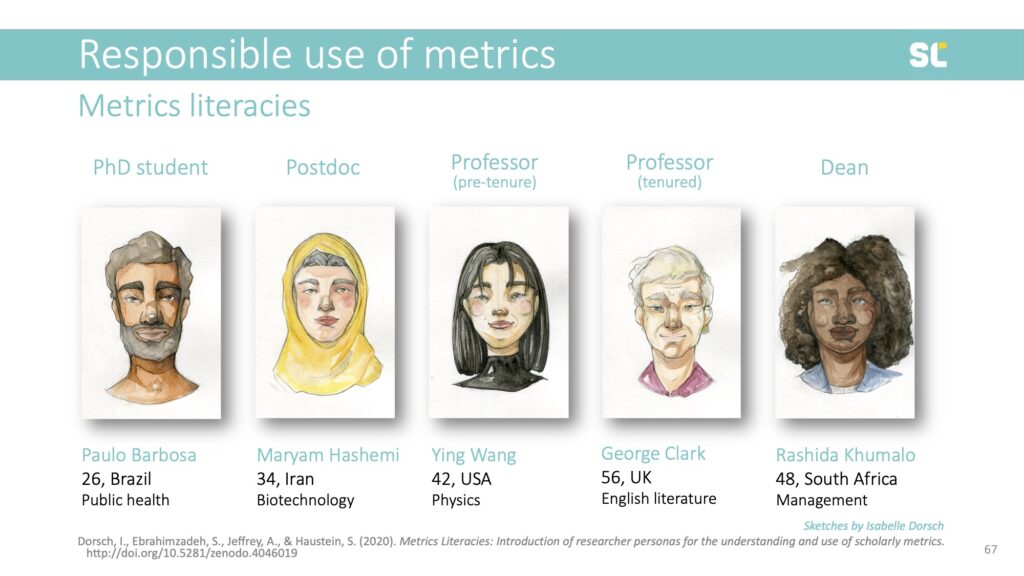
We developed personas and are employing storytelling elements to improve retention and make the videos more engaging. The personas come from different cultural and disciplinary backgrounds, and are at various career stages and roles to allow viewers to identify with them.
We believe that it needs experts in scientometrics, online education and video producers to work together to fill the current gap in metrics education in order to promote the responsible use of metrics in academia.

Conclusions
To wrap up this talk, I would like to summarize what I think are the most important opportunities and challenges of scientometrics today.
Opportunities
1. New citation databases. Current developments of new citation databases, particularly the community-led open ones, are exciting as they can potentially make research evaluation more inclusive and diverse, in terms of the researchers and types of research outputs considered.
2. Full text indexing. With more and more publications being available open access and in machine-readable format, there is a huge opportunity to further develop scientometric indicators taking into account citation context or even by complementing formal citations listed in the reference lists with mentions in the full text (as I have shown for capturing research data reuse).
3. Evidence base for changing policy. There is a huge opportunity for scientometric methods and experts to be involved in creating an evidence base to monitor science policy changes, particularly in the context of open science, open access, and research assessment reform, where policies explicitly emphasize the important role of meta research.
Challenges
1. incomplete or incorrect metadata. Incomplete or incorrect metadata may lead to the same or similar bias and lack of visibility for research from particular disciplines, countries, or publishers. I think it is up to us as the scientometric community to contribute and invest in improving this open infrastructure.
2. Data-driven indicator development. There is still a challenge of scientometric indicator development being almost exclusively data-driven, measuring what is easy to measure instead of what is desirable.
3. Misuse of metrics. Simplistic and often flawed indicators—such as the h-index, JIF, or even absolute citation counts—are all easier to obtain for the average user than more sophisticated normalized metrics and benchmarks.
4. Dominance of for-profit companies that continue to control infrastructure. Finally, I want to emphasize that despite the diversification of scientometric data providers and the growth of open infrastructure, the market of academic publishing and research evaluation is still dominated by a few for-profit companies that control academic prestige and impact.
Thank you!
Check out the Swiss Year of Scientometrics blog for recordings of Stefanie’s lecture and workshop. Visit Zenodo for all Metrics Literacies research outputs.
Swiss Year of Scientometrics Lecture: Opportunities and Challenges of Scientometrics–Part III
This blog post is the third in a four part series based on the keynote presentation by Stefanie Haustein at the Swiss Year of Scientometrics lecture and workshop series at ETH Zurich on June 7, 2023. Stay tuned for our next post, in which we’ll share opportunities and challenges of scientometrics, with a focus on the responsible use of metrics and conclusions.

Diversification of applications
I now want to talk about opportunities and challenges of scientometrics with regards to applications of scientometrics methods.
Collection management
Let’s go back to the roots of scientometrics... One of the first ever citation analyses was done by Gross and Gross, two librarians who decided which journals to subscribe to for a chemistry library.
To quote their 1927 paper in Science:
“One way to answer this question would be merely to sit down and compile a list of those journals which one considers indispensable. Such a procedure might prove eminently successful in certain cases, but it seems reasonably certain that often the result would be seasoned too much by the needs, likes and dislikes of the compiler.
In casting around for a better method of arriving at the answer, the writers decided to seek an arbitrary standard of some kind by which to measure the desirability of purchasing a particular journal. If one grants, to avoid argument, that the department is trying to train men, first, to understand the science of chemistry (including, of course, the methods and means of advancing the frontiers of the science) and, second, to be able actually to contribute to this progress, then it seems inquiry should be made into the library tools which men are using who are now doing just this.
With this purpose in mind, it was decided to tabulate the references in a single volume of The Journal of the American Chemical Society. This journal was chosen as the most representative of American chemistry. It is believed that the results of such a tabulation can be considered statistically and used with certain reservations to predict the future needs for a period, let us say, of ten years. The most recent complete volume (1926) of this journal has been chosen and the results tabulated in such a way that the relative importance of any single periodical for any five-year period can be seen. This is very important when one considers that only relatively few libraries can afford complete files of journals which have been published continuously for a century or more.”
Thus, citation analysis, but also usage analysis based on downloads, can inform collection management.
This is particularly true when it comes to canceling big deals, where having an evidence base of what content contained in bundles is actually used puts librarians in better positions when negotiating with large publishers—particularly those for-profit companies that control more than half of journal publishing.
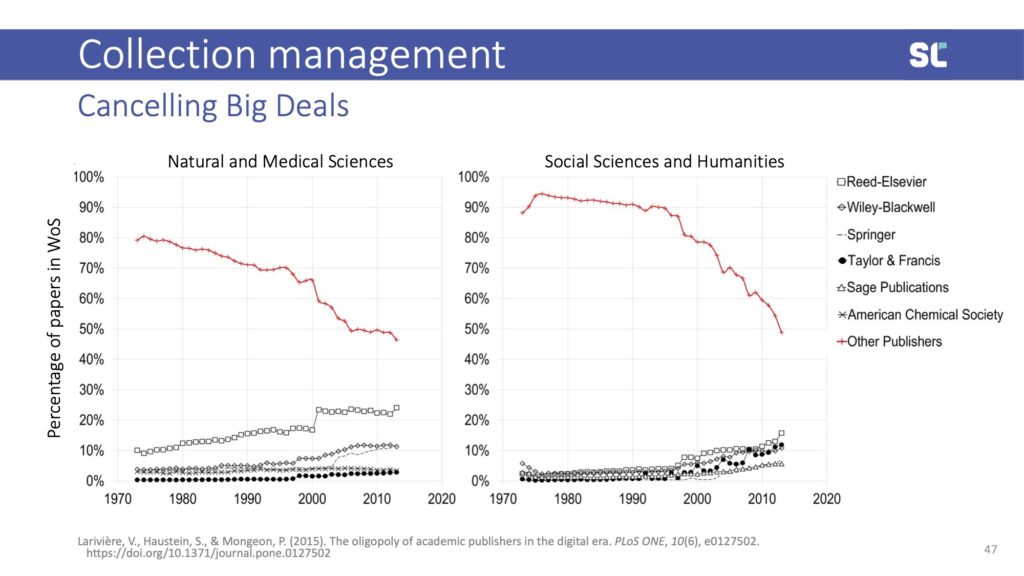
Unpaywall, and their service Unsub, can be used to investigate whether journal subscriptions are still necessary if more and more publications are available open access.
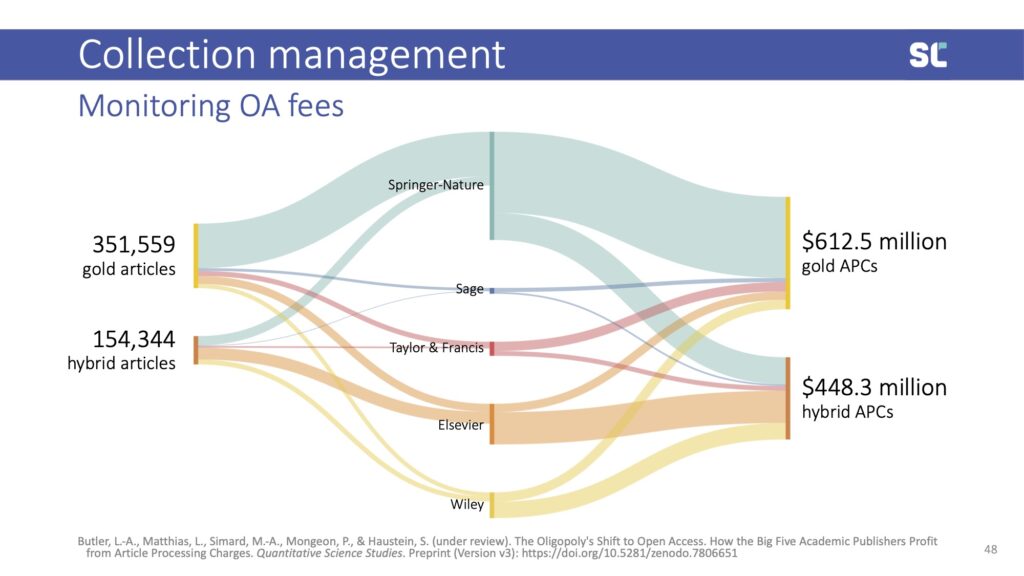
With the shift to open access, bibliometrics has now become an important tool to track the cost of article processing charges. In a 2023 preprint (under review at Quantitative Science Studies), we estimated that, globally, the academic community paid more than one billion U.S. dollars to publish open access to the five-big publishers.
Publication statistics are also used in the context of transformative agreements. For instance, it allows publishers to determine flat rate fees and justify price increases, and librarians to monitor the use of agreements.
Open science monitoring
Another new application area of scientometrics is open science monitoring. With an increasing number of open science policies, funders and policy makers are interested in tracking compliance. From open access, we know that rates of compliance with mandates are significantly higher when they are monitored.
Influenced by developments in open infrastructure, more and more dashboards are being developed to track open practices such as open access publishing and data sharing.
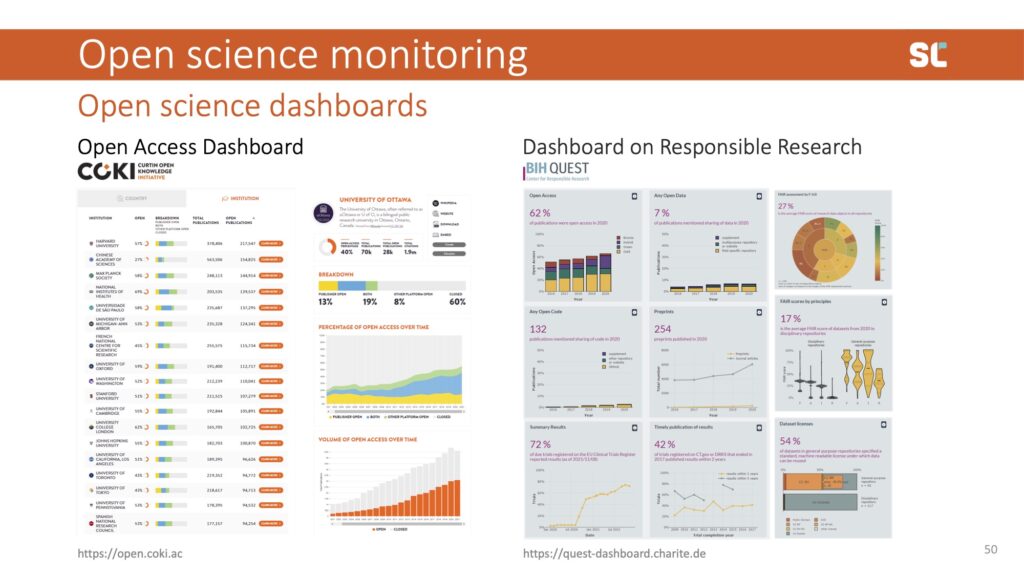
Above, I am showing examples of two dashboards: the Curtin Open Knowledge Initiative Open Access Dashboard and the Charité Berlin Dashboard on Responsible Research. There are many more examples, including the French Open Science Monitor and the European Open Science Cloud.
Most dashboards are similar to bibliometrics in the past: very data driven, tracking what's available. We are taking a different user-centred approach with the open science dashboard for biomedical institutions. Instead of including what’s easily trackable, we first conducted a Delphi study to ask the community what they consider important to open science practices and what to track in a dashboard.
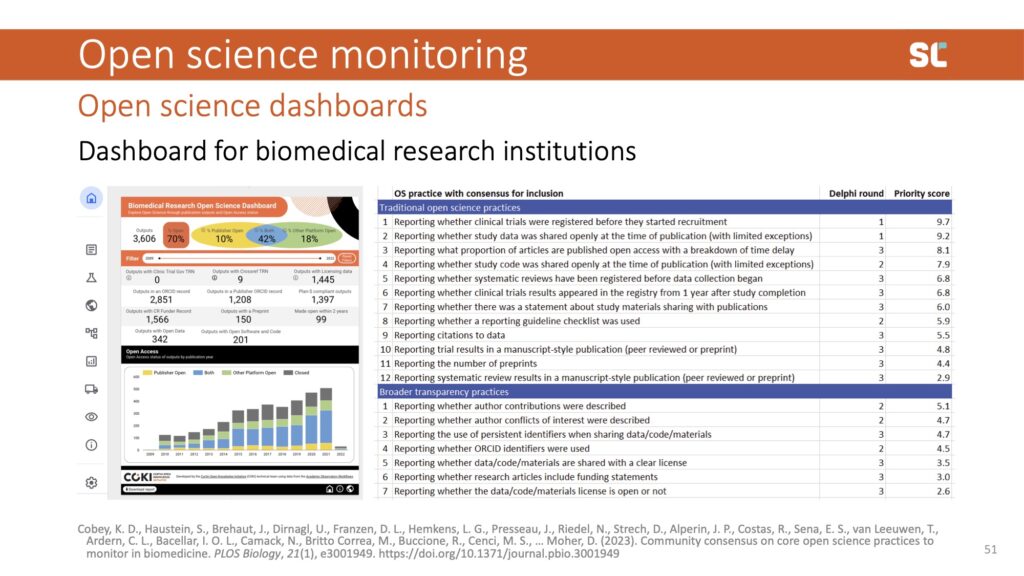
This is certainly challenging since many outputs—especially those that aren’t publications—lack the necessary metadata (e.g., institutional affiliations) to generate all indicators that the community deemed important.
Stay tuned for our next post, in which we’ll share opportunities and challenges of scientometrics, with a focus on the responsible use of metrics and conclusions. Check out the Swiss Year of Scientometrics blog for recordings of Stefanie’s lecture and workshop.
References
Gross, P.L.K, & Gross, E.M. College Libraries and Chemical Education. Science, 66, 385-389 (1927).
Swiss Year of Scientometrics Lecture: Opportunities and Challenges of Scientometrics–Part II
This blog post is the second in a four part series based on the keynote presentation by Stefanie Haustein at the Swiss Year of Scientometrics lecture and workshop series at ETH Zurich on June 7, 2023. Stay tuned for our next post, in which we’ll share opportunities and challenges of scientometrics, with a focus on the diversification of application.
Diversification of research outputs
Publications
Quantitative research evaluation and bibliometric studies in general have traditionally captured and therefore perpetuated what I would call a “biblio-monoculture,” where all that counts is publishing in English language peer-reviewed journals—which are biased towards topics and actors from the Global North, and controlled by for-profit companies that put up paywalls to limit access to literature and bibliometric data.
With the diversification of bibliometric data providers there is the hope to achieve so-called “bibliodiversity,” which values diverse contents, formats, languages, and publishing models.
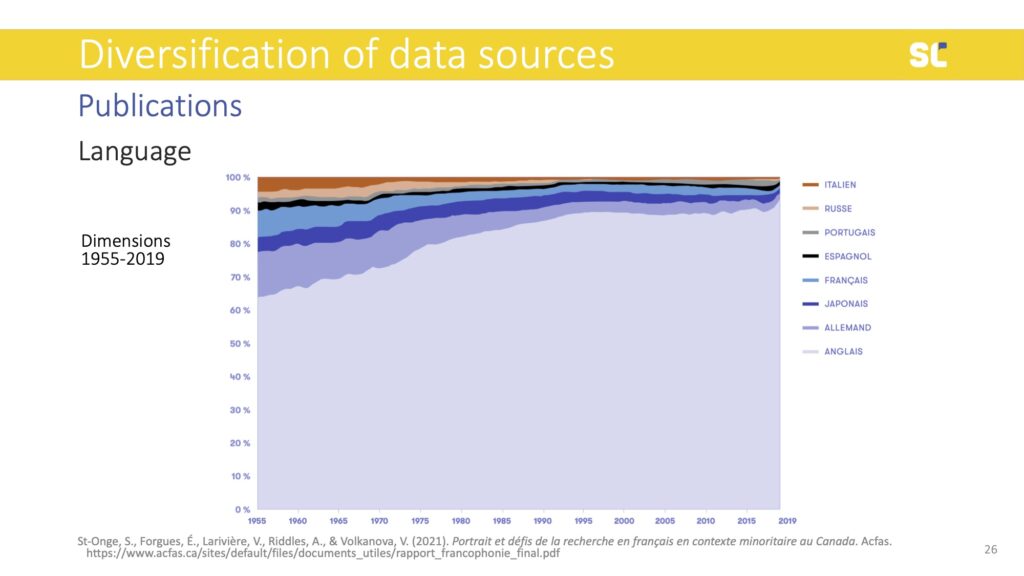
In the figure above, we see that even if databases expand, languages other than English are in steep decline. Although at different timelines, the decline of non-English content happens across all disciplines, including the humanities and social sciences where national languages and regional topics are particularly relevant.
The lack of diversity in bibliometric databases does not only show in languages but also in the representation of authors. Below you see a world map that is distorted according to the number of publications per country in Scopus. Countries like the U.S. and the U.K. were particularly overrepresented, while the Global South is extremely underrepresented.
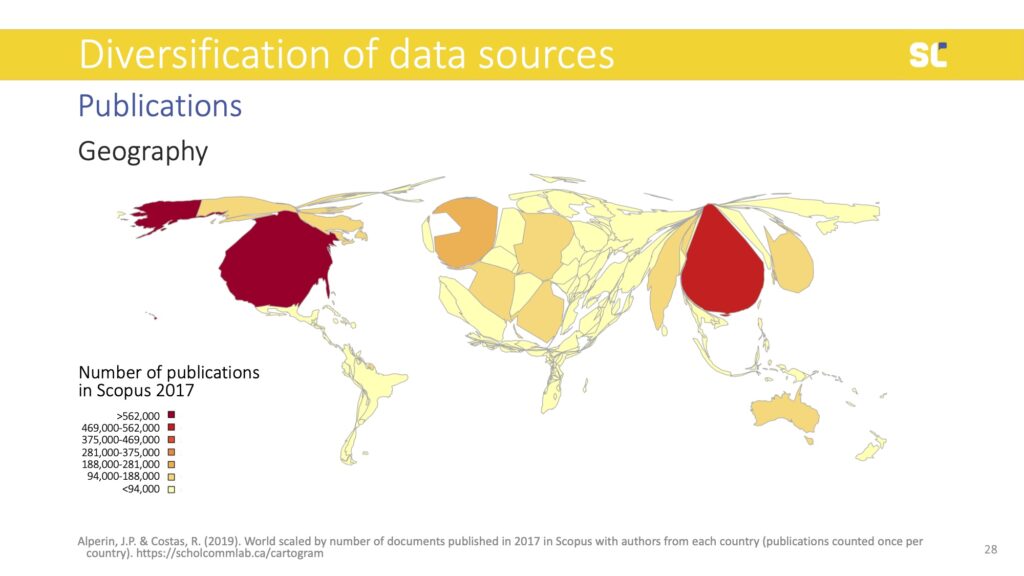
This bias in bibliometric databases has created a research culture where academic publishing that does not conform to the Western STEM model (peer-reviewed journals controlled by large publishers) is more or less invisible.
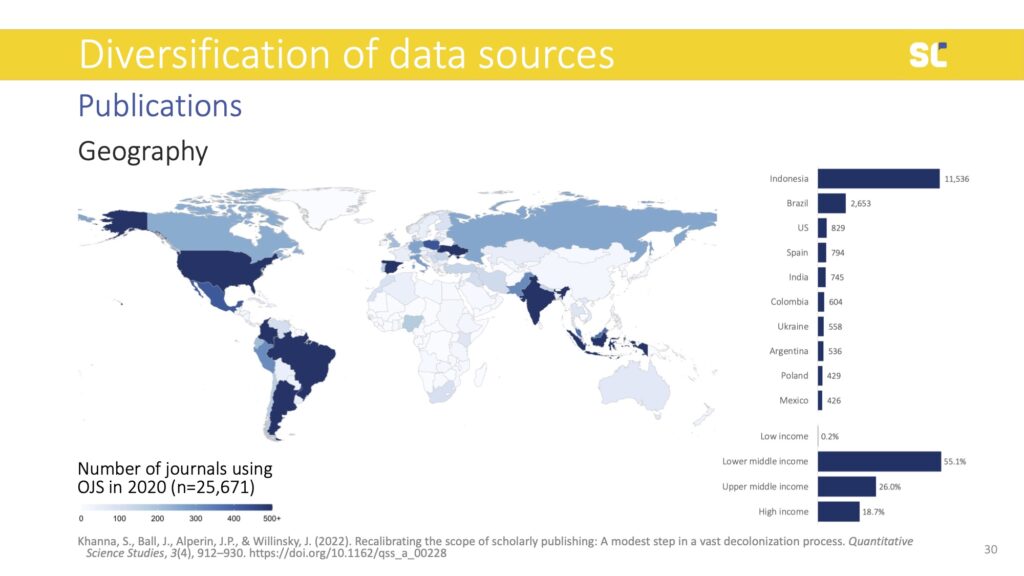
Using different, more inclusive and diverse databases means that alternative publishing models can be made visible. For example, an analysis of journals in the Directory of Open Access Journals (DOAJ) showed that there is a wealth of open access journals that exist in Asia, Latin America, and Eastern Europe that operate without paywalls for authors or readers (so called diamond Open Access journals) or with reasonable and more sustainable article processing charges than their counterparts in Western Europe and North America.
A 2022 study (published in Quantitative Science Studies) found that more than 25,000 journals that use the open-source publishing platform Open Journal Systems (OJS). 80% of the journals operated in the Global South, 84% of which were diamond Open Access. Only 1% of them were indexed in WoS and 6% in Scopus. OpenAlex indexes 64% of these journals, while Dimensions in 54%. This indicates that with these new players, scientometric studies have the potential to become more inclusive.

Data
Besides publications, data has become a valuable research output. The open science movement and an increasing number of science policies now demand or encourage researchers to share underlying data openly when they publish their papers. While there are many advantages of open data—such as increased transparency and reproducibility or fostering reuse and collaboration instead of working in silos—it’s not always easy to do so.
To quote Borgman and Bourne (2022):
“Community incentives for exchanging data are clear; incentives for individual scientists to invest in data management and distribution are less apparent. Disincentives abound, including lack of skills, lack of curatorial personnel, lack of infrastructure for managing and archiving, concerns about privacy and confidentiality of human subjects’ records, intellectual property rights, concerns about being ‘scooped,’ misuses of data, labor to assist data reusers in interpretation, cost, and more.”
I want to present some results from the Meaningful Data Counts research project that is part of the larger Make Data Count initiative that aims to address social and technical barriers to open data metrics.
In order to develop meaningful data metrics we think you need at least a minimum of two essential types of metadata: citations and information about discipline (or fields of research).
Discipline has shown to be an important factor to determine how data is reused, even what constitutes data in the first place is very different between disciplines or even smaller fields of research. For example, in STEM numerical data are very common while in the humanities text is most common.
Even if we look at our own field, what we consider as our data is bibliographic information. This is why Christine Borgman defined data in a very broad way as “entities used as evidence of phenomena for the purpose of research or scholarship.” Basically anything that is used as evidence in research constitutes data.
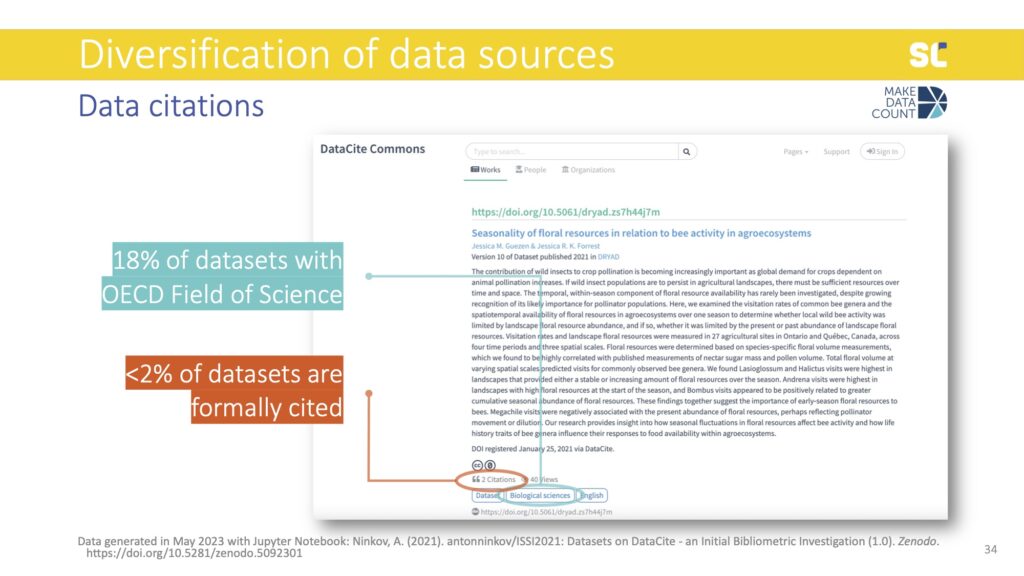

As you can see above, we are in a bit of trouble in terms of these two sources of data that we need to construct data metrics. Of the 14 million datasets in DataCite, only 18% had an Organisation for Economic Co-operation and Development (OECD) Field of Sciences and only 1.5% had been cited. If you consider datasets that have both discipline information and at least one citation, we go down to 0.6% of datasets. Therefore, we are currently lacking essential metadata for research data to construct meaningful data citation metrics.
Beyond discipline, there are plenty of other characteristics that might affect how datasets are getting reused—such as data type, file format, license, repository, etc. There needs to be more research into these to discover patterns that help create benchmarks and normalized indicators. I think what we can all agree on is that we don’t want to recreate mistakes from bibliometrics, where simple and flawed indicators—such as the Journal Impact Factor and h-index—have created adverse effects.
Tracing data
This is why we have done a large survey with almost 2,500 researchers that is stratified by academic discipline. We analyzed differences between disciplines and those items, where statistically significant differences were found between the 6 OECD Field of Sciences are indicated with an asterisk. The results that I am presenting were weighted responses to ensure disciplinary representativity.
We asked people what they would consider important to reward in the context of data work. All fields showed similar patterns for rewarding good documentation, getting credit for sharing data, and recognizing data as standalone outputs.
Significant differences between the Field of Sciences were also observed for assessing research, including the influence of your own and other people’s data as well as getting credit for reusing other people’s data. These factors were perceived as more important for people in Agricultural Science and Medical and Health Sciences than those in the Social Sciences and Humanities.
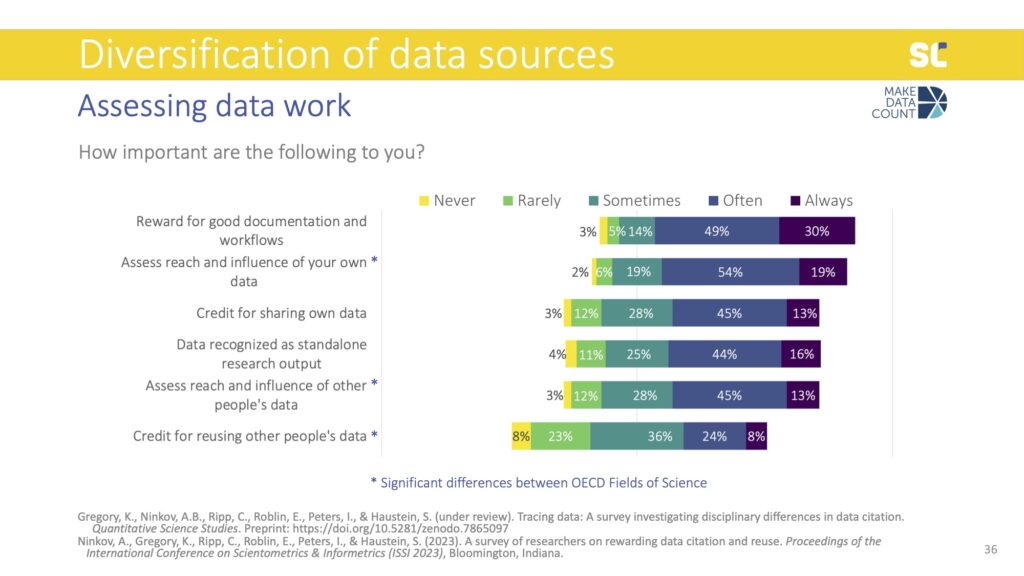
We also asked participants what would be important information to have when assessing data created by others that they might consider to reuse. The most important piece of information about data reuse was having descriptions or narratives that provide details about how the data were already used—and this was the same across all disciplines.
When assessing data created by others, people also felt it was more important to follow the number of citations, where the data was used, and who has used the data, rather than data downloads, views, and recognition outside academia. However, all of these aspects were valued differently across fields of science. For example, data citations were rated as most important in Agriculture, Engineering, and Medicine but least in the Social Sciences.
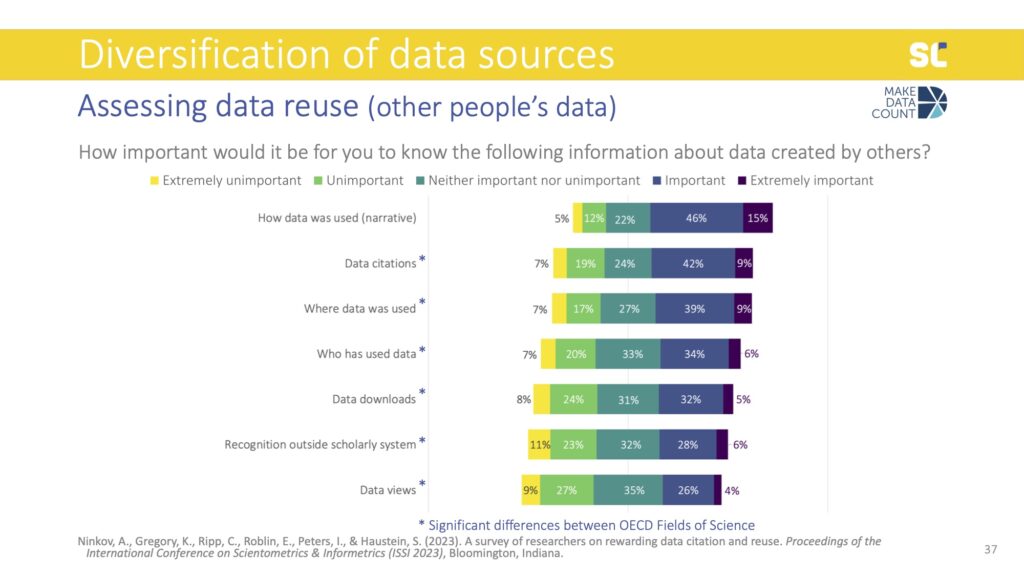
When we asked the same question but this time about their own data, all types of impact indicators were more important and data citations were considered even more essential than the narratives. Data downloads also became slightly more important than the where and who.
There were also significant differences for all items except for data citations, where all disciplines agree that this is the most important. Interestingly, the Social Sciences valued this information about their own data way more than the information about data created by others, while the opposite was true for Engineering.
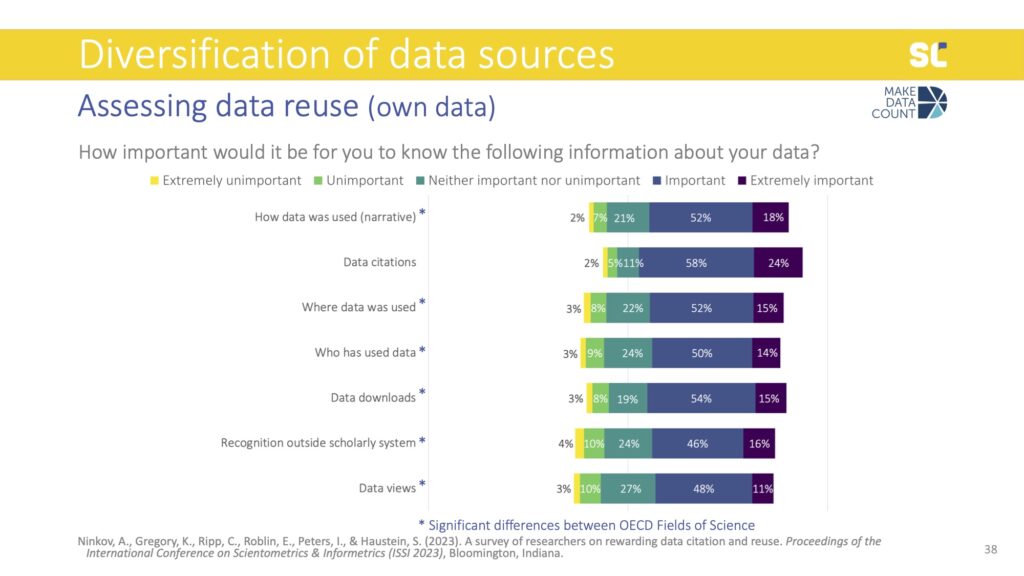
Motivations to cite data were similar to what we know about motivations to cite literature, namely showing intellectual debt (or what Robert Merton called “pellets of peer recognition”), supporting validity of claims, indicating reuse, and rewarding the data creators. Another, more altruistic reason was also to help others locate the data used in the study.
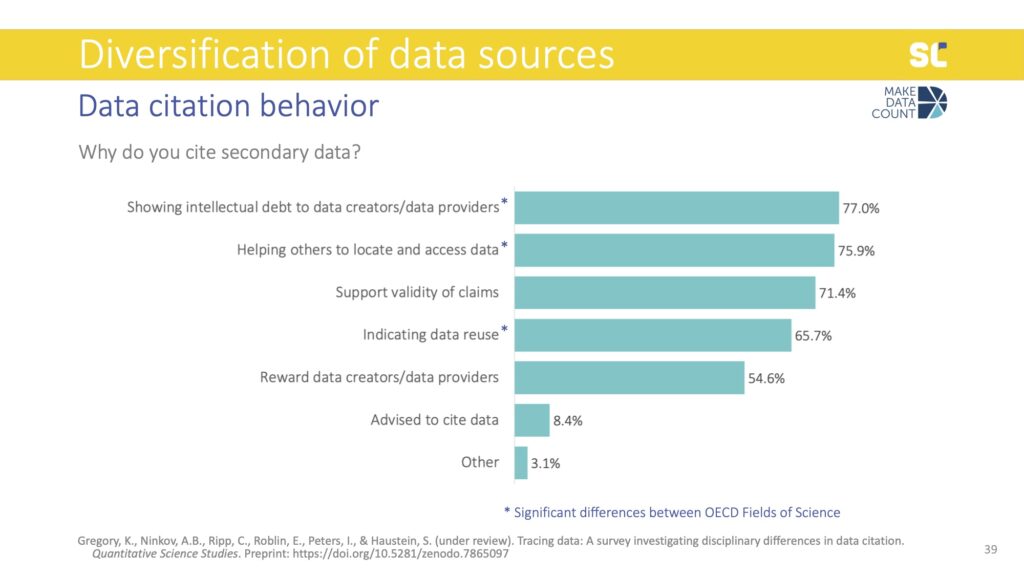
The most common way of crediting data reuse was to cite a paper analyzing the data instead of the data themselves. Citing the source of the data was also more common than actually providing a citation to the dataset directly. From a scientometric perspective this is problematic because data reuse is not directly traceable if only citing related papers.
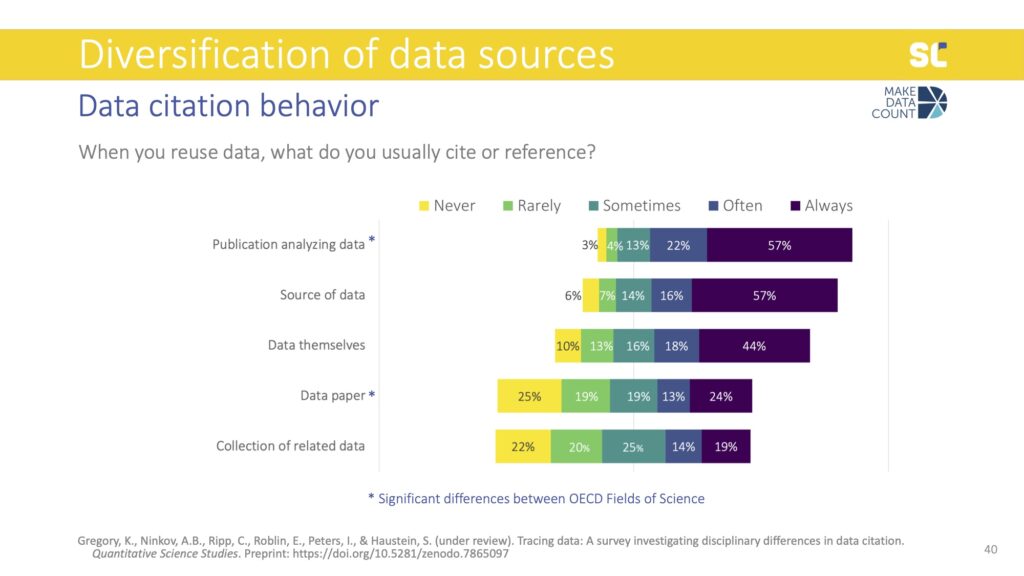
We also asked how exactly participants cite or reference data. The most common method was to list a related paper in the reference list, which is something we call indirect citations. This reflects the dominance of journal articles and metrics based on literature citations, not on data citations.
Interestingly, data citations—which means citing the data themselves in the reference list—were almost as common as indirect citations, contradicting some previous empirical findings from studies looking at how data is cited in the literature.
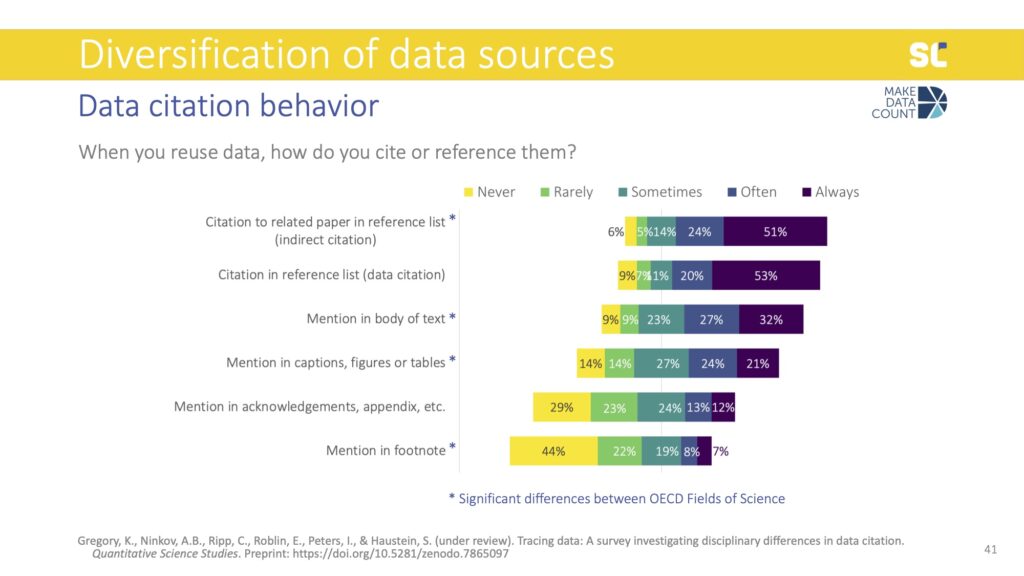
The remaining items focused on what we call data mentions: instead of formally citing data, authors mention them in different parts of the full text—particularly the body of the text but also captions, figures, or tables. This was also found in previous studies, which showed that authors may acknowledge the reuse of data in the full text of the study rather than the reference list. Through follow-up interviews with 20 researchers, we found that some think that mentioning the data in the text of caption constitutes as a citation.
Of course, from our perspective of tracking citations based on references lists, we know that this is not the case. In terms of developing data citation indicators to credit data sharing as an open science practice, a significant share of acknowledgements to data gets lost.
This is why DataCite, in collaboration with the Chan Zuckerberg Initiative, recently received funding from the Wellcome Trust to extract data mentions from the full text of academic literature.
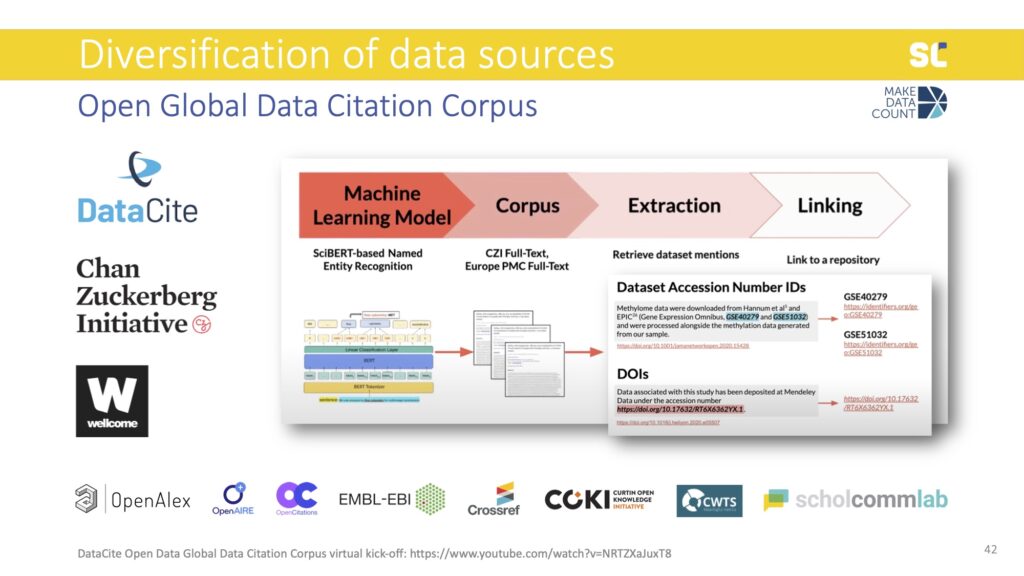
They are developing a machine learning algorithm that is able to extract dataset mentions from full text to fill this gap. The Open Global Data Citation Corpus will thus contain both data citations—as DataCite was previously capturing—as well as data mentions. In addition to DOIs, it will also expand to track dataset accession number IDs, which are more common than DOIs in some fields (mostly in biomedical research).
The Open Global Data Citation Corpus will be an important step towards the development of data metrics in order to reward open science practices, acknowledge the diversification, and value research outputs beyond publications.
Stay tuned for our next post, in which we’ll share opportunities and challenges of scientometrics, with a focus on the diversification of applications. Check out the Swiss Year of Scientometrics blog for recordings of Stefanie’s lecture and workshop, and Zenodo for all Meaningful Data Counts research outputs.
References
Borgman, C. L., & Bourne, P. (2022). Why It Takes a Village to Manage and Share Data. Harvard Data Science Review, 4(3).
Swiss Year of Scientometrics Lecture: Opportunities and Challenges of Scientometrics–Part I
This blog post is the first in a four part series based on the keynote presentation by Stefanie Haustein at the Swiss Year of Scientometrics lecture and workshop series at ETH Zurich on June 7, 2023. Stay tuned for our next post, in which we’ll share opportunities and challenges of scientometrics, with a focus on the diversification of research outputs.

Thank you to David Johann and Annette Guignard at ETH Library for inviting me to open this lecture series for the Swiss Year of Scientometrics.
In the hope of inspiring some interesting discussions, my goal is to provide you with a very broad overview of a number of opportunities and challenges of scientometrics, with a focus on the diversification of data sources and areas of application.
I will talk about how the data sources for bibliometric analysis are becoming more diverse, both from the perspective of data providers and the types of research outputs considered.
With respect to new areas where scientometrics can generate an evidence base, I will briefly address library collection management, open science monitoring as well as the responsible use of scholarly metrics.
I’ll conclude the talk with some reflections on what I consider the main opportunities and challenges of scientometrics today.
Diversification of data providers
The number of companies providing bibliometric data has really exploded in recent years. Some of you might still remember the times when the Web of Science was the only option for citation analysis.
While closed infrastructure and proprietary data sources were the norm for decades, we now see an explosion of open infrastructure developments. In my opinion, a particularly exciting development is the flip from closed to open infrastructure—when OurResearch took over Microsoft Academic Graph (MAG) and turned it into OpenAlex.
The open players—OpenAlex, Crossref and DataCite—are committed to a set of guidelines called the Principles of Open Scholarly Infrastructure (POSI). These guidelines prescribe, for example, that the infrastructures should be steered collectively by the community, not by lobbyists, and that revenue generation is mission oriented and based on services, not on selling data. In fact, all data and software of these organizations should be open and not patentable.
OpenAlex
When Microsoft decided to cease operations, the non-profit OurResearch with support from a $4.5 million grant from Arcadia decided to further develop MAG and turn it into open infrastructure that is accessible via API, a data dump, and (since very recently) a user interface.
Apart from it being open and free to reuse for any purposes, OpenAlex’s unique selling point is certainly its coverage and size, making it a great potential source for more inclusive bibliometric analyses.
Approaching its metadata from a bibliometric perspective, OpenAlex provides the basic building blocks for the analysis of research outputs and impact. OpenAlex is a heterogeneous directed graph linking research outputs, called Works, as well as information about who published them, where they were published, their research area, and even who funded it.
Since OpenAlex was only recently launched, we don’t know much yet about the feasibility for scientometric analyses. So I want to explore two crucial metadata elements in a bit more detail: institutional affiliations and disciplines.
Author addresses on academic publications provide us with information about their institution. In this example, you see that this author was affiliated with Simon Fraser University in Canada:
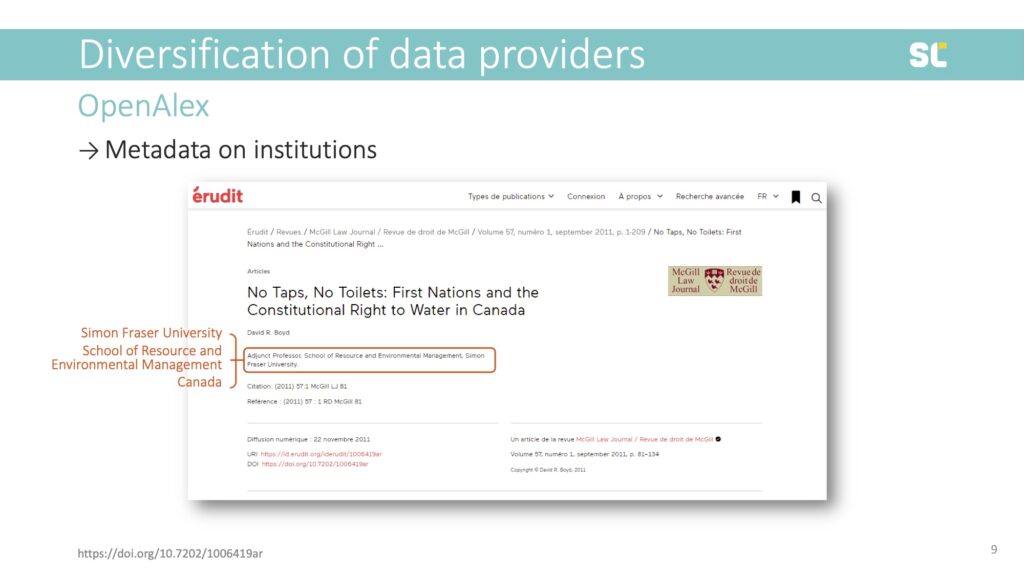
These elements form the basic building blocks for an institutional analysis on the meso and macro levels of bibliometric analysis, providing us with the university, department, and country of the author.
OpenAlex uses an algorithm to link institutional information to the Persistent Identifier from the Research Organization Registry (ROR ID), combining metadata from Crossref, PubMed, ROR, Microsoft Academic Graph, and publisher websites. Through this enriching process, every institution in OpenAlex is supposed to have a ROR ID.
In this example, however, we see that OpenAlex does not have any institutional metadata even though the original publication did:
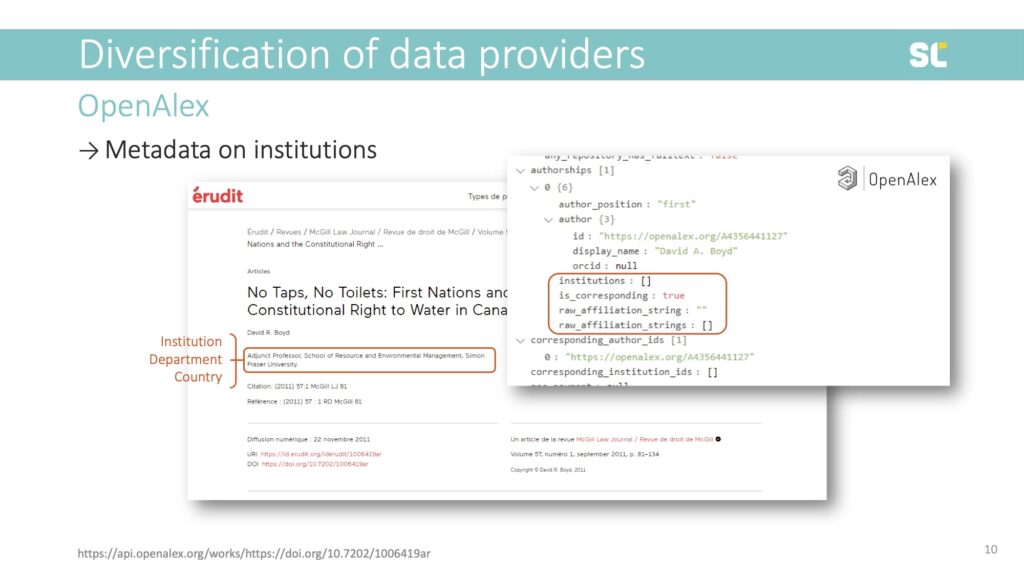

By the way, the same is true for this paper in Dimensions, which likely means that the publisher did not provide institutional information to Crossref.
So what does this mean bibliometrically? While this article could be tracked via DOI, author name, or journal, it would not be included in any analysis that relies on country or institution, either as a method of retrieval, or unit of analysis. This paper would neither count as an output for Canada nor Simon Fraser University because the metadata is missing.
My colleague Simon van Bellen at the Canadian journal platform Érudit analyzed the presence of RORs in OpenAlex for all papers published in Canadian social sciences and humanities journals from 2012 to 2021. He found that 63% of papers were retrievable via ROR ID, which means that over one third of papers did not have an institutional address.
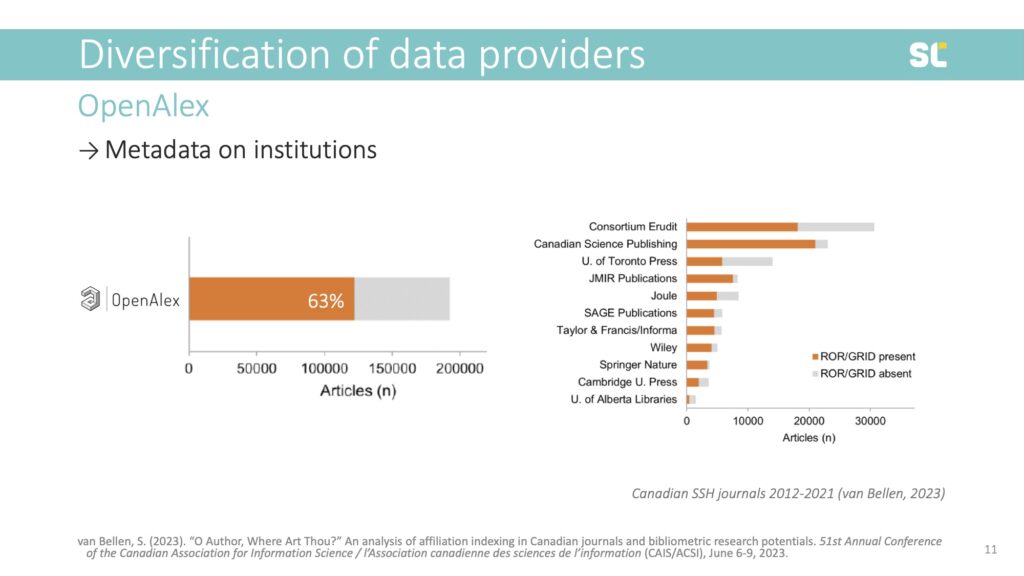
What is even more alarming is the huge differences between publishers. For example, while institutional metadata was present in 90% of journals published by Springer-Nature, the non-profit Consortium Erudit had institutional metadata in only 59% of their published papers.
This means that the metadata that publishers provide affects the visibility of these papers in bibliometric analyses using OpenAlex or Dimensions. So even if these larger more inclusive databases now cover more outputs, they don’t necessarily translate into more and more diverse outputs on the institutional or country level.
Disciplinary metadata is particularly important in scientometric analyses for benchmarking and normalizing field-specific publication and citation practices.
OpenAlex uses Wikidata concepts for subject indexing. Concepts are assigned automatically based on title, abstract and journal, conference or book title. Concepts are hierarchical with five levels for a total of 65,000 concepts.
This classification system is a modified version from Microsoft Academic’s fields of study, which has been criticized by Sven Hug and colleagues (2017) who concluded that it would not make a good base for bibliometric indicators due to the dynamic number of fields and inconsistent hierarchies.
Although they are made to look like a true hierarchical classification system with an astonishing number of classes, concepts do NOT fulfill the properties of a regular classification system. For example, in the slide below, you see the first two hierarchies and that many of these have two parent classes.
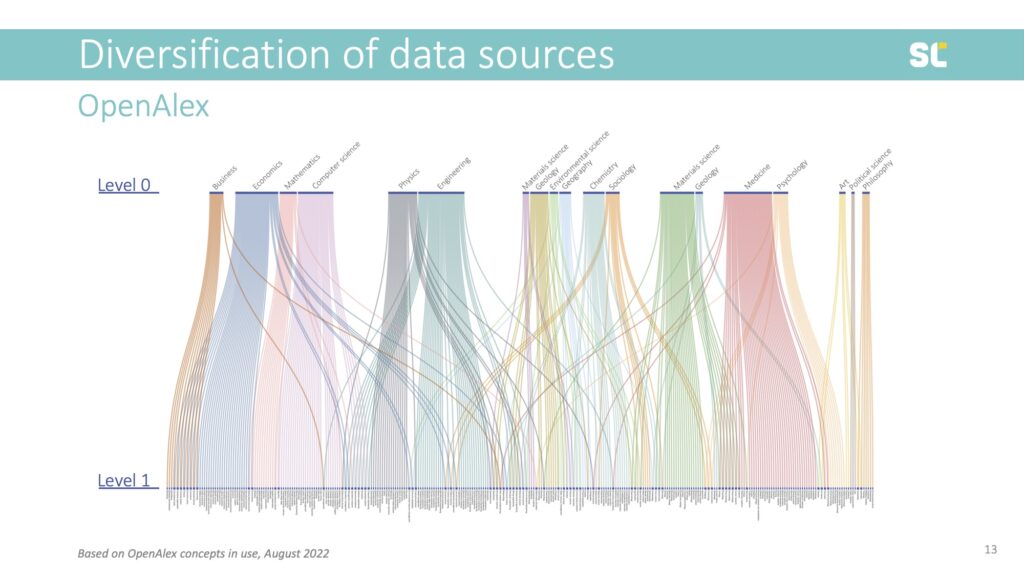
Moreover, the quality of some index terms is questionable and so are some of the hierarchical relationships. For instance, Literature L1 (under L0 Art) has 391 level 2 classes, including geographical classes (Italian literature, Scottish literature), classes about literary movements (modernism, absurdism) or literary criticism (reception theory), genres (detective fiction, travel writing) as well as concepts that certainly shouldn’t be found on the third hierarchy level, or not even be classes at all—such as individual Shakespeare plays like Hamlet, Star Trek, or Harry Potter[1].
Another severe issue is also that there is no vocabulary control in the OpenAlex concepts. For example, homonyms are not distinguished. As shown in the slide below, this book review published in the Sixteenth Century Journal and this conference paper published in a Lecture Notes in Computer Science book series are tagged with the same concept.
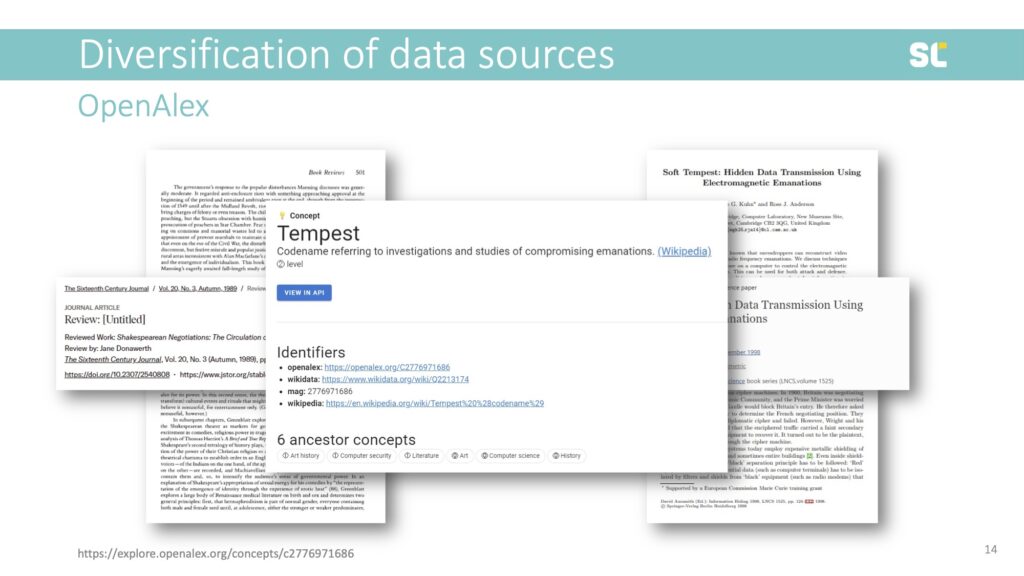
“Tempest” is a Level 2 term (third level of hierarchy) and a subordinate class of Literature, Art History, and Computer Security. The Tempest is a play by Shakespeare but also an acronym for “Telecommunications Electronics Materials Protected from Emanating Spurious Transmissions,” which is the US National Security Agency’s specification for protecting against data theft through the interception of electromagnetic radiation.
Another example are these two papers, one published in a journal called Agricultural and Food Chemistry and one in IEEE Signal Processing Letters.
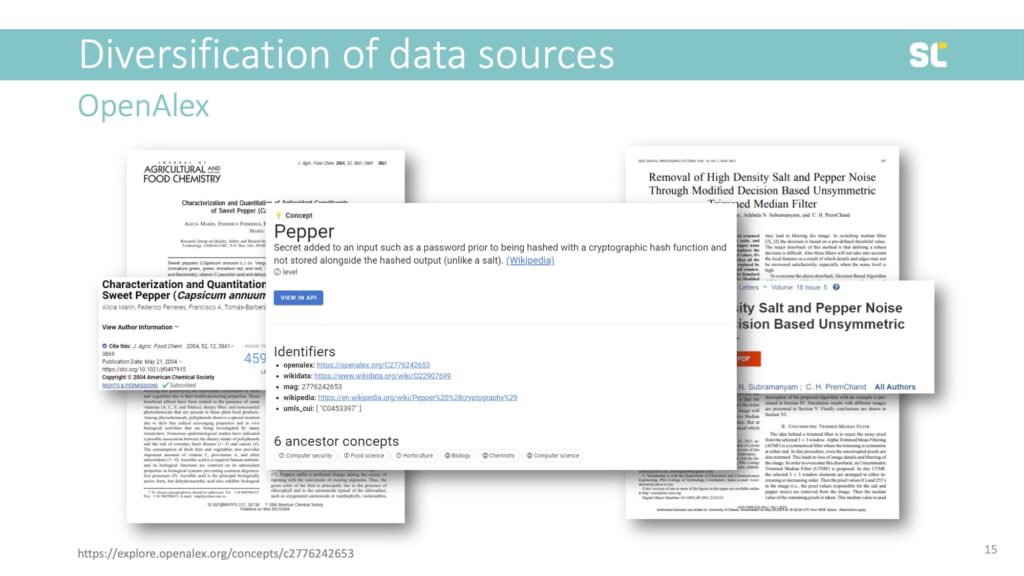
“Pepper” is a Level 2 class, with three different parent classes: Food Science, Horticulture, and Computer Security. This shows that the concepts in OpenAlex are highly problematic and should at this point not be used for bibliometric analyses.
However, I want to emphasize the recent launch of OpenAlex and that in my opinion it provides a huge opportunity to build an open, community owned system that can be more inclusive if metadata quality and completeness are addressed. I think that we need to invest in this open infrastructure and contribute to improving quality as a community.
Elsevier
I now want to talk about Elsevier, who has been a bibliometric data provider through Scopus for several decades. But instead of talking about Scopus as such, I want to highlight how it has become an information analytics business that has been collecting metadata of scholarly outputs for almost 150 years.
I also want to highlight the aspect of Elsevier being a highly profitable business. As Sarah Lamdan highlights in her recent book Data Cartels, “Elsevier focuses on crunching academic researchers’ data in new ways to make money off the research process.”
Elsevier is part of the RELX Group, representing the Scientific, Technical, and Medical segment. What is clear is that it is no longer just a publisher, it’s a data analytics company. Databases, tools, and electronic reference generate almost as much revenue as publishing.
But it’s because of the wealth of information and decades of electronic and online data collection that Elsevier has become so powerful. They have built an incredible data analytics pipeline by collecting data and acquiring tools and platforms that capture the entire research cycle and research evaluation process (see below).
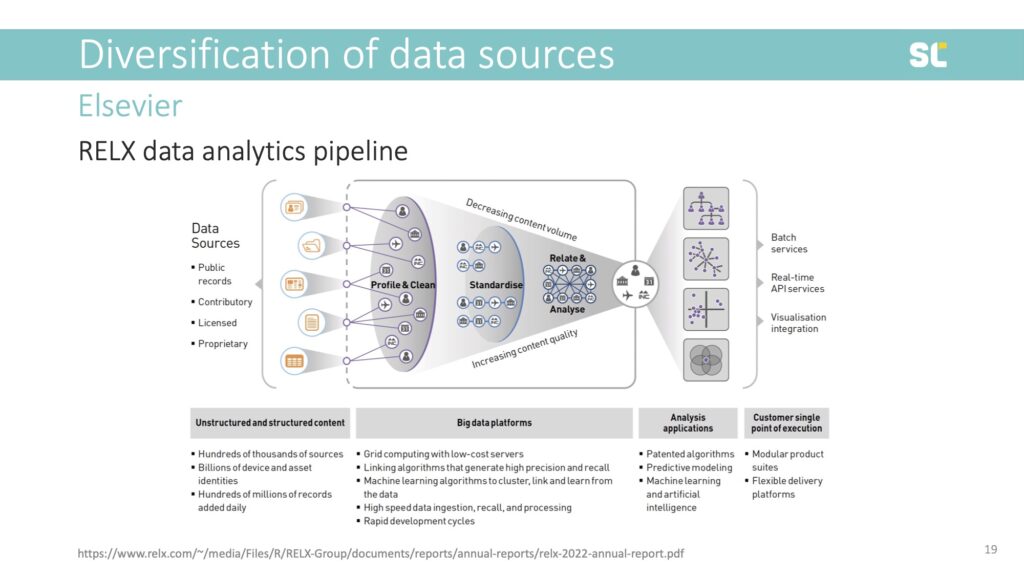
This would be incredible infrastructure for the academic community but because Elsevier is a private company and not a community organization working for the public good, it does what it’s supposed to do: “monetize the entire knowledge production cycle” (Lamdan, 2023, p. 53) and sell raw data from content created by the academic community and data about people who access this content, back to its customers in a plethora of services.
Last year alone, Elsevier created 2.9 billion pounds in revenue and 1.1 billion pounds profit. In the last 10 years, as shown below, their profit was 9.1 billion pounds. That’s almost 12 billion Swiss franc or 11.3 billion euros.
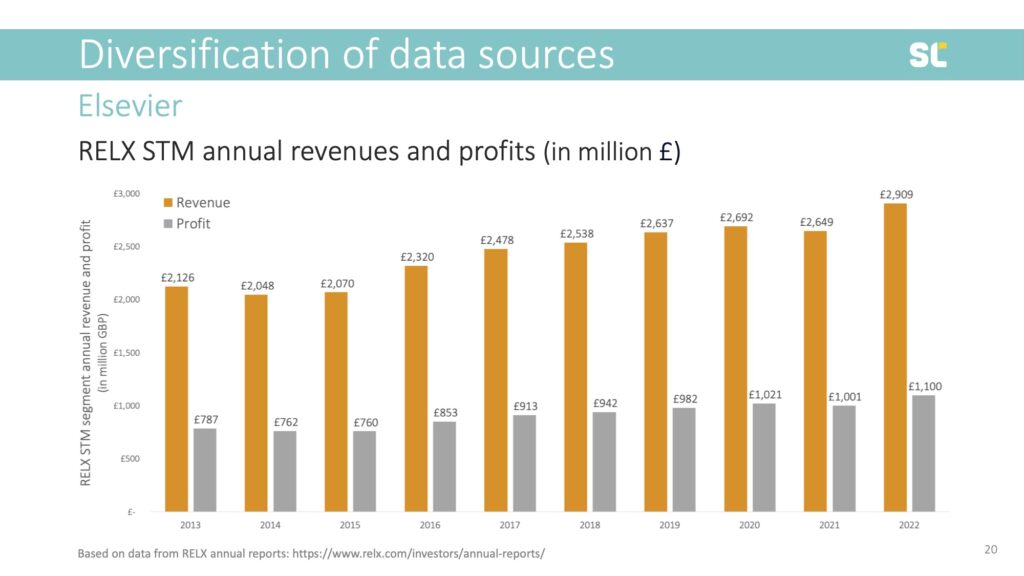
Elsevier’s profit margin is very stable, at 38% annually, because most of their content is provided for free, be it authors’ manuscripts, peer review, or the digital traces left by those accessing the content. This means that for every 1,000 euro spent by the scholarly community on article processing charges, journals or database access, 380 euro leaves the system to pay shareholders—people that were smart enough to invest in RELX stock.
Looking ahead, Elsevier is expecting to grow and further increase profits.
From a scientometric perspective, Elsevier controls and maintains a wealth of rich, connected metadata about the entire research lifecycle. For the academic community, this comes at unsustainable costs. It makes one think what could be possible if just a fraction of these amounts would go into supporting open infrastructure, both for publishing and scientometric analyses.
Stay tuned for our next post, in which we’ll share opportunities and challenges of scientometrics, with a focus on the diversification of research outputs. Check out the Swiss Year of Scientometrics blog for recordings of Stefanie’s lecture and workshop.
References
Hug, S.E., Ochsner, M. & Brändle, M.P. Citation analysis with microsoft academic. Scientometrics 111, 371–378 (2017).
Lamden, S. Data Cartels: The Companies That Control and Monopolize Our Information, Stanford University Press (2023).
van Bellen, S. "O Author, Where Art Thou?" An Analysis of Affiliation Indexing in Canadian Journals and Bibliometric Research Potential. CAIS-ACSI 2023, Canada. Zenodo (2023).
Footnotes
[1] Special thanks to University of Ottawa ÉSIS students Alexandra Auger, Karinne M., and Caroline Plante for analyzing the OpenAlex concepts Literature class in detail.
ScholCommLab at ACFAS 2023
By Chantal Ripp, with editorial support by Olivia Aguiar
The 90th annual ACFAS conference was held at the Université de Montréal, HEC Montréal, and Polytechnique Montréal from May 8 to May 12, 2023. Over the years, the ACFAS Congress has become the largest multidisciplinary gathering of Francophone scholars, attracting participants from around the world.
Under the theme "100 years of knowledge for a sustainable world," this edition offered a rich scientific program to highlight the place of research and knowledge in our societies, and their contributions to the common good.
Seven lab members—Juan Pablo, Rémi Toupin, Anton Ninkov, Chantal Ripp, Leigh-Ann Butler, Marc-André Simard, and Stefanie Haustein—attended or presented sessions at the "Perspectives sur la publication et les revues savantes : évolution du libre accès, des pratiques d’évaluation et des nouvelles possibilités de recherche (Perspectives on publishing and scholarly journals: Evolution of Open Access, review practices and new research opportunities)" colloquium.
-

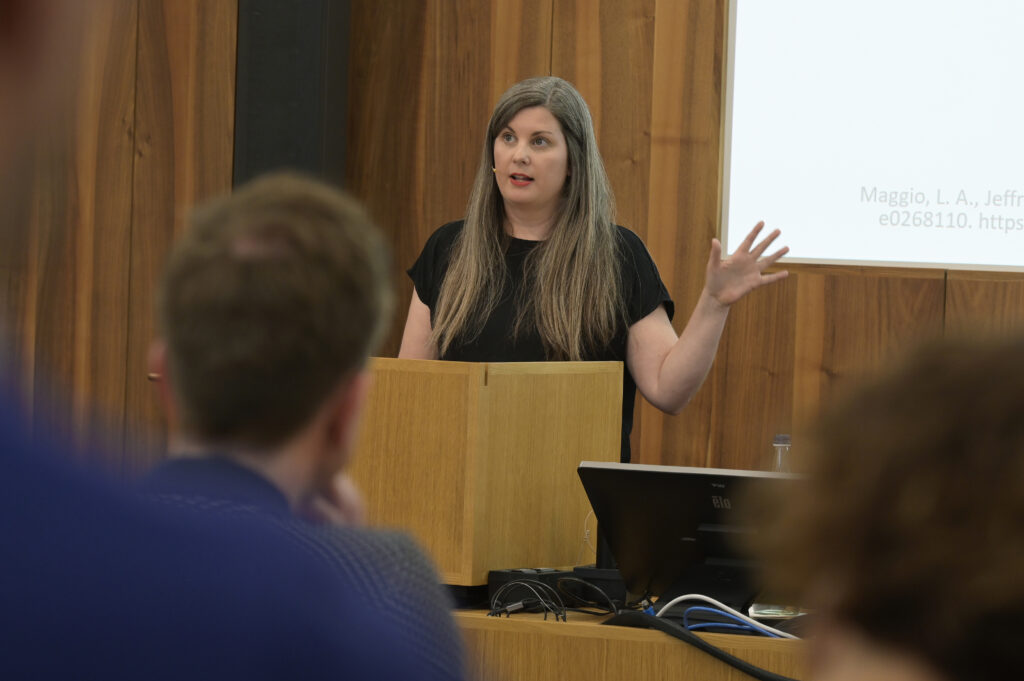
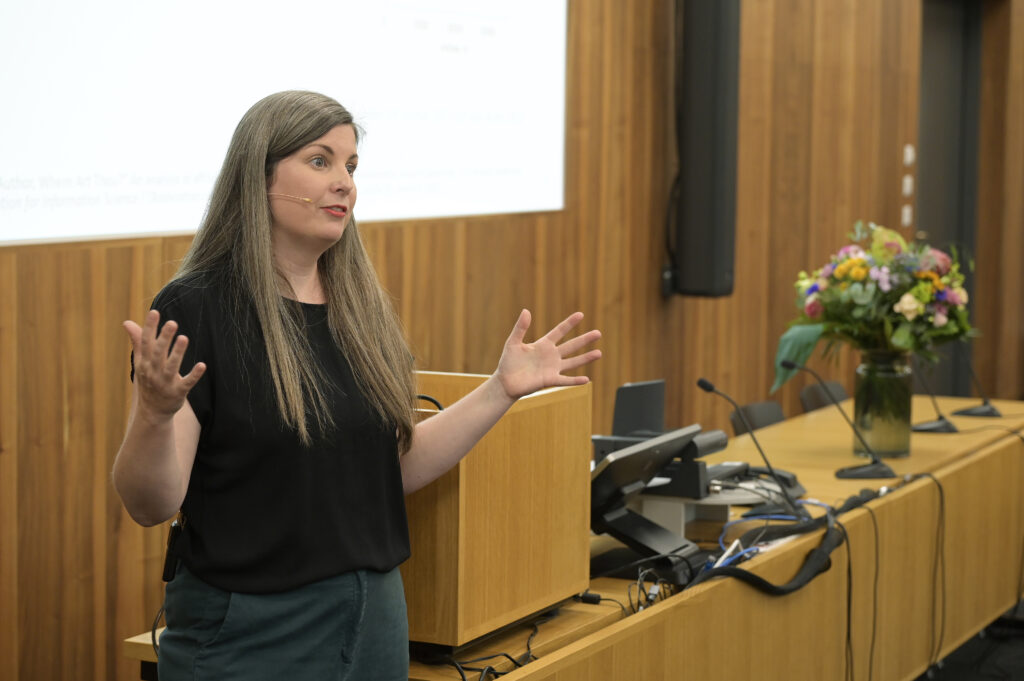
Leigh-Ann Butler (left) and Stefanie Haustein (right) co-present on “New models, same problems: How the big five publishers' open access business models perpetuate inequalities in scholarly communication” at ACFAS 2023 | Photo by Chantal Ripp -
Marc-André Simard presents on “International comparison of open access journals and their use of article processing fees” at ACFAS 2023 | Photo by Chantal Ripp. -
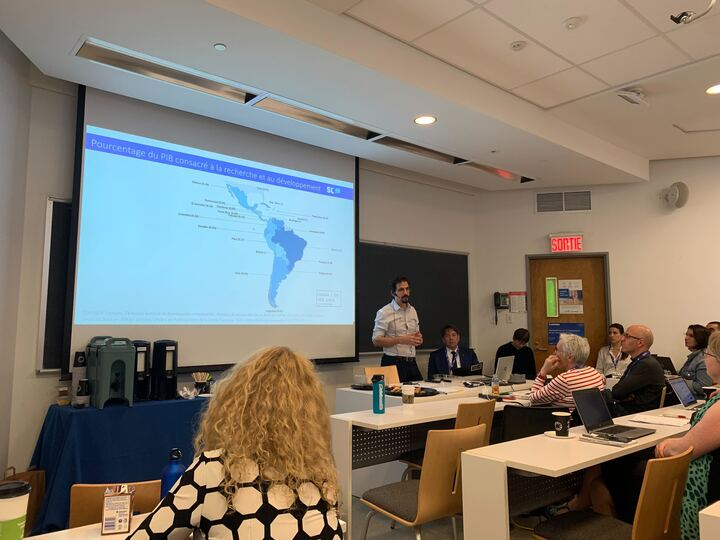

Juan Pablo Alperin presents on “Publication incentives in Latin America” at ACFAS 2023 | Photo by Chantal Ripp
The two-day colloquium explored the evolution of scholarly publishing, new opportunities offered by digital research infrastructures, as well as challenges and opportunities faced by Quebec and Canadian journals as they transition to open access. The participation of many lab members at the conference reflects the lab’s promotion and dissemination of scholarly communication research in French and English.
In addition to the captivating sessions at ACFAS, members of the lab were also excited to (re)connect in-person and chat over dinner. "Having a virtual space to collaborate with colleagues is great, but getting the opportunity to spend time in person is worthwhile, especially as so many of the lab members are spread across Canada. As a new PhD student, it's a great way to feel that sense of camaraderie," says Chantal Ripp. "It was my first ACFAS event and I am looking forward to attending again!"

For more on the conference, check out the conference website and program. The University of Ottawa will host the 91st annual ACFAS conference in 2024.
Introducing VOICES: Understanding equity and inclusion in open science
By Olivia Aguiar and Alice Fleerackers
During the pandemic, more research was shared openly, more preprints were posted, and we saw an explosion in the public communication of science, particularly in mainstream media. In the long-term, these changes have the potential to foster more open, diverse, and inclusive approaches to research and bolster our capacity to face present and future societal challenges. Yet, whether that potential will become a reality remains an open question.
To help realize this potential, the ScholCommLab launched a project to better understand the Value of Openness, Inclusion, Communication, and Engagement for Science in a Post-Pandemic World (VOICES). The project brings together a transnational team of scholars (from Brazil, Canada, Germany, and the UK) with complementary expertise in open science, scholarly and science communication, and research impact. Through four interconnected subprojects, it aims to explore the new interplay between researchers, policymakers, science communicators, and the public that has affected science and society during the COVID-19 pandemic and beyond.
In this post, VOICES scholars share their hopes for the project and reflect on key learnings from the official kickoff meeting that took place in Hamburg, Germany this November. The meeting raised important questions about the role of research in society, what it means to change “science culture,” and how to foster meaningful collaboration within interdisciplinary teams.

How ‘open’ is open science?
Open Science (OS) advocates have long argued that making science more open will lead to increased participation from diverse stakeholders, particularly those who have traditionally been excluded from science. But, in practice, this may not always be the case. VOICES aims to provide much needed empirical evidence into whether the uptake of OS practices seen during the pandemic has made science more inclusive or diverse.
“I have learned from my research and from working on the ground that ‘openness’ doesn’t automatically lead to accessibility and inclusion,” says Natascha Chtena, a postdoctoral fellow at the ScholCommLab and research coordinator of VOICES. “I hope our work will help push conversations and actions toward an open science that is better anchored in society and better able to serve diverse communities around the world.”
Isabella Peters, a professor of Web Science at ZBW Leibniz-Information Center for Economics and Kiel University, Germany and a principal investigator (PI) on VOICES, agrees: “I hope that we will better understand what are the true effects of open science and whether the goals of open science - that are often only mentioned implicitly, such as greater inclusion - are actually reached.”

To investigate these assumptions, VOICES will address three main questions situated before and during the pandemic:
(1) How is the value of open science discussed and positioned?
(2) Who adopted or contributed to open science practices and how?
(3) How has the relationship between research and the public been affected by the opening of research?
The work will identify OS practices that can foster inclusive, sustainable scholarly systems, greater public engagement with research, and a more resilient society. “I am looking forward to seeing how this project manages to put forward a vision and understanding of what is needed for open, inclusive, and equitable science by drawing on international perspectives that span at least three continents,” says Juan Pablo Alperin, an associate professor in Simon Fraser University’s Publishing Program, co-director of the ScholCommLab, and lead PI of VOICES.
Fostering collaboration within a transnational research team

VOICES is funded by a Trans-Atlantic Platform Recovery, Renewal, and Resilience in a Post-Pandemic World award. This highly competitive funding scheme aims to support scholars from both sides of the Atlantic to collaborate on research investigating the “effects of the pandemic on all aspects of health, social, economic, political, and cultural life.”
The VOICES kickoff workshop in Hamburg revealed just how enriching it can be to collaborate across disciplinary and geographic borders. “Working with people from different countries not only gathers different fields and scholar’s background perspectives but also adds cultural flavour to research as an enriching and pleasant process,” says Germana Barata, a science communication researcher at the University of Campinas (Unicamp) and PI of the Brazilian VOICES team.
According to Natascha, establishing synergy between projects is not a passive process. “Collaboration doesn’t just happen because you bring a group of people together. It takes effort, intentionality, and a true commitment to the idea that we are better together,” she says. “As we continue our collaboration, the thing that excites me most is identifying synergies and complementarities between our different sub-projects, learning from each others' unique perspectives, and finding meaningful ways to contribute to each other’s work.”

The Hamburg meeting was an important step towards establishing those meaningful collaborations. Over two days, each national team presented and shared research progress, relevant literature, and action items for the other teams. All four teams also worked on developing a framework which integrates inclusion, participation, and engagement within OS, that extends beyond the academic community. The workshop sparked vibrant and challenging conversations, both within and across the national teams.
“I was excited to discuss our ideas for our bibliometric data and to see how it relates to our project goals and the overall developments in the field of open science,” says Isabelle Dorsch, a postdoctoral fellow on the project supporting the team at the ZBW – Leibniz Information Centre for Economics in Kiel. She enjoyed the opportunity to share different knowledge and viewpoints on how the projects may “click” together.
While sharing these different views was enriching, it also raised difficult questions. “The workshop helped me recognize the importance of context: to notice how my experiences and background shape my understanding of issues like ‘equity’ or ‘participation,” says Alice Fleerackers, PhD candidate in the ScholCommLab and researcher on the Canadian team. “It revealed how difficult—but also how rewarding—it can be to collaborate across contexts.”

Towards a more inclusive open science
As the pandemic subsides, many will be faced with a choice of going back to “business as usual” or continuing to adopt open, diverse, and inclusive practices. For the VOICES team, finding ways to help scholars, journalists, and science communicators work towards a more inclusive open science in the aftermath of COVID-19 will be a priority.
There’s still a long way to go before achieving that goal, but the research team is hopeful for the future. “I hope this project will encourage researchers to think more critically about open science—to ask who we are including when we make our work ‘publicly available’ and which voices we might be leaving out,” says Alice.
To stay up to date with the VOICES project and other news from the lab, sign up for the ScholCommLab newsletter.
Journalists reporting on the COVID-19 pandemic relied on research that had yet to be peer reviewed
By Alice Fleerackers (Simon Fraser University) and Lauren A Maggio (Uniformed Services University of the Health Science)

(Shutterstock)
A story on gender inequity in scientific research industries. A deep dive into the daily rhythms of the immune system. A look at vaccine effectiveness for COVID-19 variants. These are a few examples of news stories based on preprints — research studies that haven’t been formally vetted by the scientific community.
Journalists have historically been discouraged from reporting on preprints because of fears that the findings could be exaggerated, inaccurate or flat-out wrong. But our new research suggests that the COVID-19 pandemic may have changed things by pushing preprint-based journalism into the mainstream.
While this new normal offers important benefits for journalists and their audiences, it also comes with risks and challenges that deserve our attention.
Peer review and the pandemic
Traditionally, studies must be read and critiqued by at least two independent experts before they can be published in a scientific journal — a process known as “peer review.”
This isn’t the case with preprints, which are posted online almost immediately, without formal review. This immediacy has made preprints a valuable resource for scientists tackling the COVID-19 pandemic.
The lack of formal review makes preprints a faster way to communicate science, albeit a potentially riskier approach. While peer review isn’t perfect, it can help scientists identify errors in data or more clearly communicate their findings.
Studies suggest that most preprints stand up well to the scrutiny of peer review. Still, in some cases, findings can change in important ways between the time a study is posted as a preprint and the time it is published in a peer-reviewed journal, which can be on average more than 100 days.
A ‘paradigm shift’ in science journalism
As researchers of journalism and science communication, we’ve been keeping a close eye on media coverage of preprints since the onset of the pandemic. In one study, we found that a wide range of media outlets reported on COVID-19 preprints, including major outlets like The New York Times and The Guardian.
Unfortunately, many of these outlets failed to mention that these studies were preprints, leaving audiences unaware that the science they were reading hadn’t been peer reviewed.
We dug deeper into how and why journalists use preprints. Through in-depth interviews, we asked health and science journalists about the strategies they used to find, verify and communicate about preprints and whether they planned to report on them after COVID-19.
Our peer-reviewed, published study found that preprints have become an important information source for many journalists, and one that some plan to keep using post-pandemic. Journalists reported actively seeking out these unreviewed studies by visiting online servers (websites where scientists post preprints) or by monitoring social media.
Although a few journalists were unsure if they would continue using preprints, others said these studies had created “a complete paradigm shift” in science journalism.
A careful equation
Journalists told us that they valued preprints because they were more timely than peer reviewed studies, which are often published months after scientists conduct the research. As one freelancer we interviewed put it: “When people are dying, you gotta get things going a little bit.”
Journalists also appreciated that preprints are free to access and use, while many peer-reviewed journal articles are not.
Journalists balanced these benefits against the potential risks for their audiences. Many expressed a high level of skepticism about unreviewed studies, voicing concerns about the potential to spread misinformation.

(Unsplash/The Climate Reality Project)
Some journalists provided examples of issues that had become “extremely muddied” by preprints, such as whether to keep schools open during the pandemic.
Many journalists said they felt it was important to label preprints as “preprints” in their stories or mention that the research had not been peer reviewed. At the same time, they admitted that their audience probably wouldn’t understand what the words “preprint” or “peer review” mean.
In addition, verifying preprints appeared to be a real challenge for journalists, even for those with advanced science education. Many told us that they leaned heavily on interviews with experts to vet findings, with some journalists organizing what they described as their “own peer review.”
Other journalists simply relied on their intuition or “gut” instinct, especially when deadlines loomed or when experts were unavailable.
Supporting journalists to communicate science
Recently, media organizations have started publishing resources and tip sheets for reporting on preprints. While these resources are an important first step, our findings suggest that more needs to be done, especially if preprint-based journalism is indeed here to stay.
Whether it’s through providing specialized training, updating journalism school curricula or revising existing professional guidelines, we need to support journalists in verifying and communicating about preprints effectively and ethically. The quality of our news depends on it.![]()
Alice Fleerackers, PhD Student, Interdisciplinary Studies, Simon Fraser University and Lauren A Maggio, Professor, Uniformed Services University of the Health Sciences

This article is republished from The Conversation under a Creative Commons license. Read the original article.
Three questions with… Olivia!
Our lab is growing! In our Three Questions series, we’re profiling each of our members and the amazing work they’re doing.
Our latest post features Olivia Aguiar, a lab manager at the ScholCommLab and a doctoral student at Simon Fraser University’s Injury Prevention and Mobility Lab (IPML). In this Q&A, she tells us about creating science comics, interviewing "super seniors," setting boundaries, and more.

Q#1 What are you working on at the lab?
I’m so excited to be joining the ScholCommLab as a Lab Manager. My tasks include coordinating lab and project team meetings; updating the blog, website, and newsletter; collaborating on grant proposals; and supporting the lab directors in communication and outreach. I look forward to meeting the entire team and embarking on this new journey!
Q#2 Tell us about a recent paper, presentation, or project you’re proud of.
I recently completed my certificate in Biomedical Visualization and Communication (BMVC) at The University of British Columbia. During the program, I learned that I love to illustrate scientific and health comics. Currently, I'm working on a short comic about concussions in ice hockey. My goal is to finish the final version by the end of Summer 2022. Stay tuned on social media (@livsaguiar) to see the result!

Q#3 What’s the best (or worst) piece of advice you’ve ever received?
I interviewed “Super Seniors”—individuals aged 85 years or older who have never been diagnosed with a major medical concern (e.g. heart disease, cancer, dementia)—for the Healthy Aging Study at BC Cancer. My colleague and I summarized their advice on how to live a long, healthy life. My personal favourites include: “Kindness doesn’t cost anything,” “Consider life as an adventure,” and “Do good for others but also look after yourself.” Their words provide me with perspective and inspiration—especially during uncertain and challenging times.
As for professional advice, the best and worst I've received: “If you want something done right, do it yourself.” This mantra got me through university. It helped me build resilience, solve problems, and achieve success in volunteer and leadership positions. But this advice also led to many, many bouts of stress and burnout. I learned (the hard way) that sometimes there is a better person for the job. Now, I know there is a difference between pushing out of your comfort zone and pushing beyond your limits. I embrace team work and collaboration in the classroom and work place. It is brave to ask for help!
Find Olivia on Twitter at @livsaguiar
A bridge to access research: Reflections on the Community Scholars Program developmental evaluation
By Andrea Whiteley, with editorial support by Alice Fleerackers
What happens when you provide the community with free access to academic research?
Simon Fraser University (SFU)’s Community Scholars Program (CSP) is a unique initiative that aims to do just that. Established in 2016 in collaboration with the United Way of the Lower Mainland and Mindset Social Innovation Foundation, the program connects more than 500 people working in nonprofits and community organizations across BC with the latest scholarly literature, providing free access as well as research training and support.
While the program has grown considerably since its launch, it hasn’t always been clear how participating in CSP benefits community members. Curious to find out more, I met with program librarian Heather De Forest during the final stages of my PhD on Open Access and the Public Good. With support from the United Way, Mitacs, and Dr. Juan Pablo Alperin at the ScholCommLab, my casual curiosity quickly developed into a fully-fledged postdoctoral research project evaluating the success of the CSP.
In this blog post, I summarize key findings and recommendations from the final report (available to download here), drawing on survey data, group interviews, and one-on-one conversations with more than 100 participating Community Scholars.

1. Wide-ranging impact
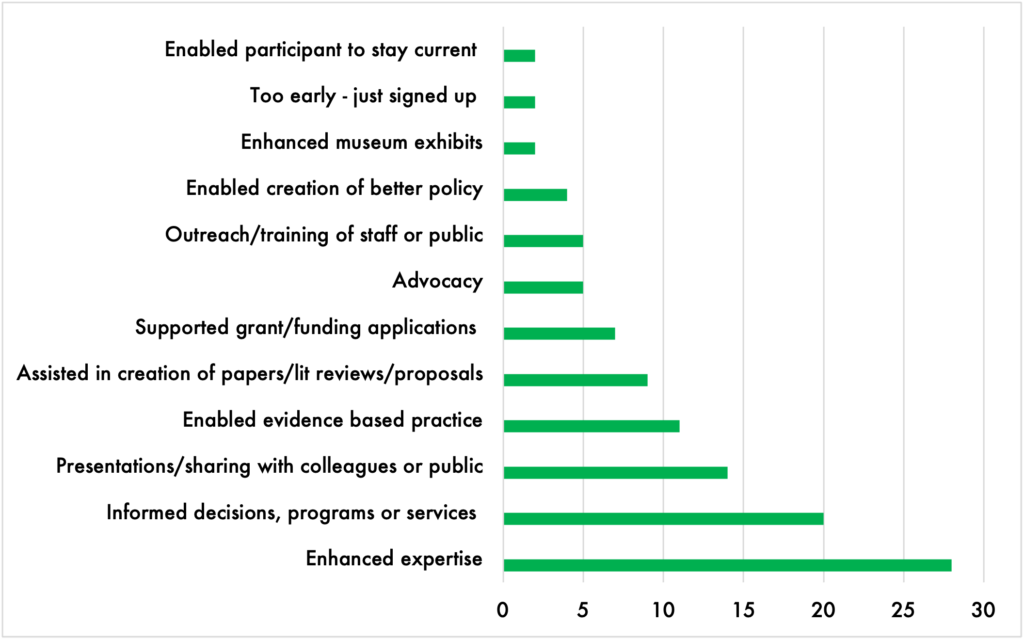
Overwhelmingly, interviews and survey responses with program participants confirmed that the CSP was having both direct and indirect impact on their lives, their organizations, and the constituencies they serve. Scholars reported appreciating personal educational impacts, like improved presentation skills or program development abilities, which they had developed through CSP’s professional workshops and training sessions. They also reported benefiting from access to the research itself:
Last year I was asked to participate in an international course as a subject matter expert on child rights. Although that is my area of work and focus I was anxious to ensure that I ‘knew enough,’ was current and was able to provide knowledge to a broad range of participants. Through CSP I was able to deepen my knowledge and gain different perspectives. Not only did this improve my confidence, it also ensured that I was able to coach 100+ participants.
The impact of the CSP on individual scholars also reverberated throughout the communities and in the nonprofit organizations where Community Scholars worked. By contributing to the professional development of employees, organizations were able to tap into a research base for successful grant funding applications, participate in legal challenges, enhance their ability to deliver evidence-based programs, create research and new knowledge in their fields, and more.
2. Professional development and lifelong learning
Although over 60% of Community Scholars in the study had graduate level education (and by extension some research background), many expressed an interest in receiving further research-related education. The CSP program currently provides participants with various opportunities for training, which one Scholar called opportunities for “group study” and “self-study.”
In surveys and interviews, many Community Scholars said that these opportunities filled a gap in their professional development, allowing them to enhance their understanding of their field, supporting in-house research, and motivating them to learn more about issues in their profession. In a couple of interviews, Community Scholars pointed to a need for adult education programs, like the CSP, that could support people who do not have formal educational credentials in entering post-secondary studies. At least one participant said that participating in the program encouraged them to pursue further education.
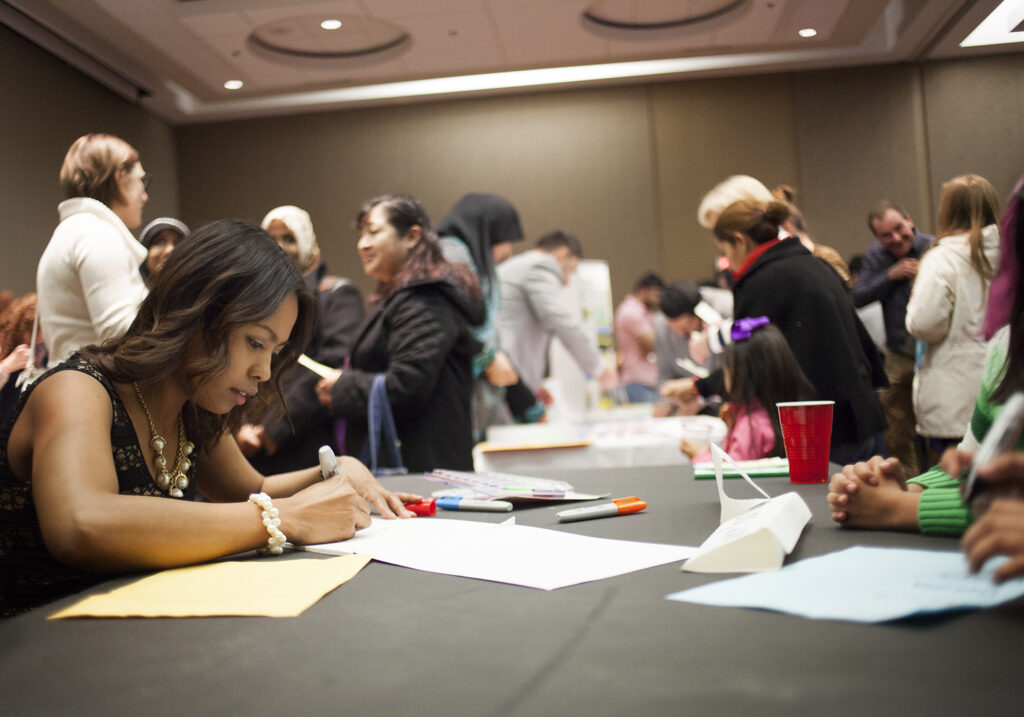

3. The power of communities of practice
The CSP brings together individuals working in the same field through journal club meetings and workshops, offered in person or online. Community Scholars who participated in journal club meetings—where participants choose research articles to read and discuss together—were very positive about their experiences and enjoyed getting to know others working in their field. Sitting in on these sessions, I was struck by how similar they were to graduate student seminars, but where each “student” possessed vast lived experience surrounding the issues under discussion.
Although many Community Scholars expressed interest in participating in more of these “communities of practice” activities, fitting networking and learning opportunities into their busy lives was a challenge. Almost two thirds (60%) of survey respondents said that time was a barrier to full participation.
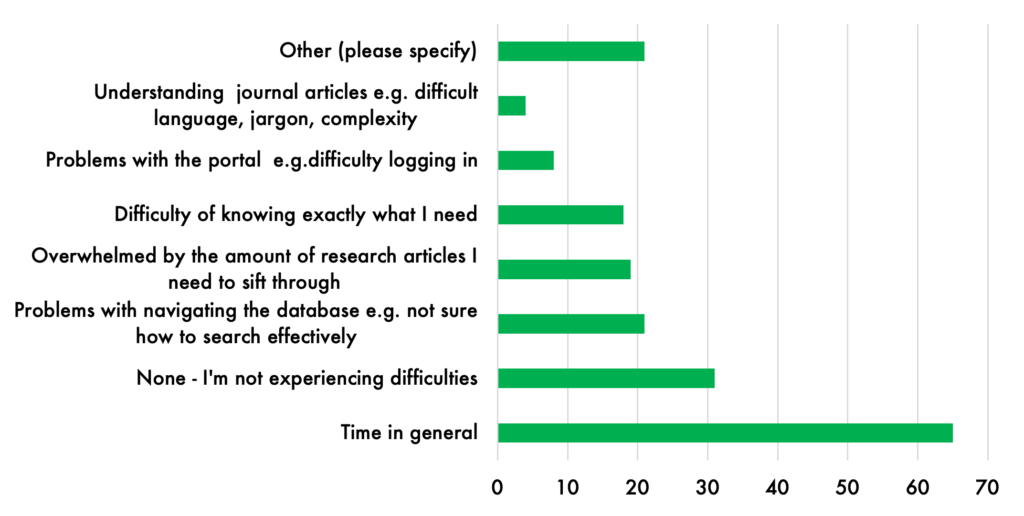
4. Librarians as essential supports
Feedback from Community Scholars confirmed that the program could not have achieved any level of success without the administrative, intellectual, and relationship management provided by the CSP librarians. During interviews, participants indicated that they received valuable assistance from the CSP librarians and were thankful to have them as a resource. Especially for Community Scholars located in remote locations, CSP librarians were the human faces of the online research portal. It was clear that librarians play an instrumental role in the transformation of community members into “Community Scholars.”
The CSP librarian in my area has been consistently warm in reaching out to offer support, both online and through workshops and the journal club.
Importantly, CSP librarians wear many hats and possess diverse skills. Employed at university libraries, they have extensive research skills. But they must also provide other kinds of support to help community members access research through the portal. Librarians demonstrated a range of innovative approaches for recruiting, training, and educating Community Scholars, including using local libraries to host information sessions, offering curated training sessions for organizations, and enlisting colleagues to present workshops on research-related topics.
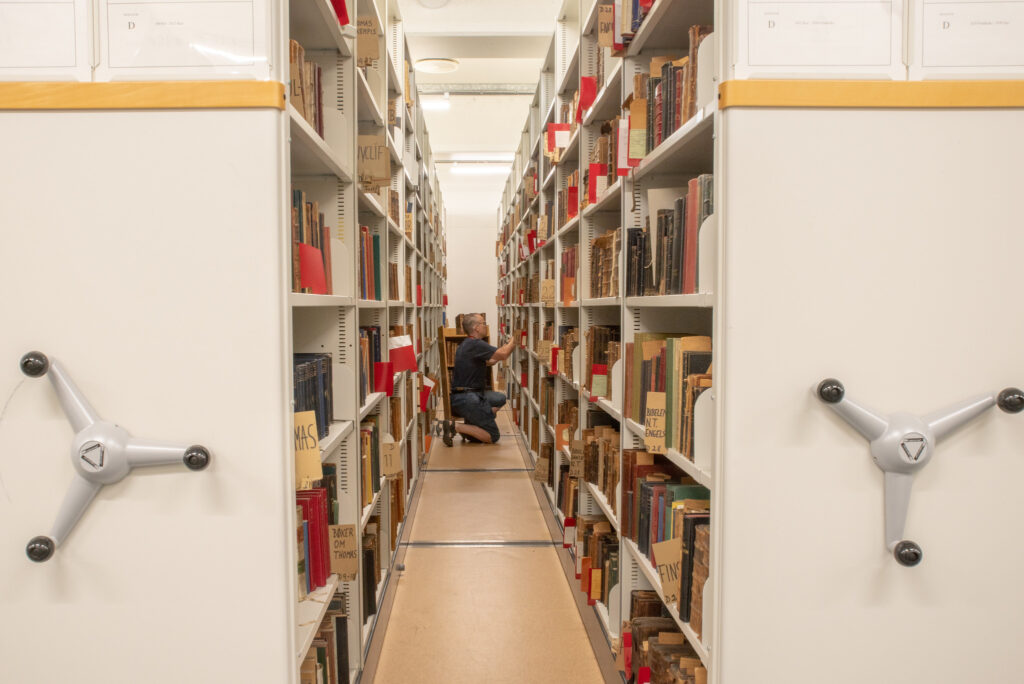

5. A bridge between disconnected worlds
Finally, my research revealed that the CSP acts as a bridge between disconnected worlds, bringing together traditional, for-profit scholarly publishing models with a more “public good”-oriented approach to knowledge access. As the success of the program depends on the willingness of scholarly publishers to allow community access to scholarship, its very existence is a living compromise within a publishing ecosystem where access to research has become a hotly contested topic. During my evaluation, I encountered countless compelling examples of the research impact made possible by bringing these seemingly conflicting realities together.
It's all bridge building, I actually consider myself a bridge builder. And right now, it's moving ... between indigenous and non-indigenous communit[ies], but there are so many bridges between communit[ies] and institutions, all of these bridges that we've been talking about. And I think Community Scholars is a flash point for that.
In a similar vein, my research revealed the potential of the program to act as a liaison between academics and their greater communities. The CSP encourages the flow of knowledge among professional scholars, community scholars, frontline service deliverers, community and nonprofit staff, policymakers, and citizens. Still, some noted that this flow of knowledge was often unidirectional. These comments point to an opportunity for the research community as a whole to develop avenues for two-way communication between scholars and their professional counterparts working in the field, and ultimately, better mobilize the knowledge created beyond university walls.

Recommendations for a path forward
While overall support for the CSP was high, several broad recommendations emerged from my evaluation that could improve current initiatives or guide the program into new territory. For example, some Community Scholars expressed interest in the idea of a “research hub” where they could be linked up with graduate students, postdocs, or other researchers to tackle specific research needs within their organizations. As one of the CSP librarians, Kate Shuttleworth, put it:
What most of our nonprofits really want is a researcher. Access is great, [the] portal is great, but we need someone to do the work. And that’s the primary thing that would help them.
The CSP already has a relationship with Simon Fraser University’s Community Engaged Research Initiative (CERI), making this organization an obvious candidate for further promoting community-based research initiatives and engagement with Community Scholars.
The evaluation also uncovered a need to develop research literacy more broadly, as nonprofit organizations are increasingly challenged by funders to implement evidence-based practices, evaluate program impacts, and understand the research landscape of their respective disciplines. The CSP could provide Scholars with guidance in this area by showing them how to conduct literature reviews, write grant proposals, or find funding opportunities. Large funders, like the United Way, could support such initiatives by clearly communicating expectations for impact reporting and evidence-based programming from a funder perspective.
One grant proposal required statistics and sources to support our request for funding. Using data and journal sources found through your portal helped us secure $255,000 in funding.
Finally, a key motivation behind this evaluation was to find ways to sustain the program in its next phase of development. Providing access to research is a cornerstone of the CSP and, although Community Scholars clearly value that access, few if any participants I spoke with felt they could afford to pay for it without the program’s support. Although my evaluation did not include a targeted analysis of possible funding and administrative supports for the CSP, my consultation with users and discussions with program leaders indicate this is an area that urgently requires further investigation. For example, the CSP could explore offering an adult education component that would create funding opportunities, particularly if expanded to include a formal “research certificate.” The CSP would also qualify for funding initiatives aimed at improving knowledge mobilization and research outreach to the public, especially in remote Indigenous and Northern BC communities.

Final thoughts: Fostering connection and culture
Above all, this evaluation made it clear that, while the CSP provides access to research, Scholars also value the “culture of scholarship” that accompanies that access. The CSP fosters a culture of collegiality and sharing, allowing participants to engage with scholarly research while also taking into account the unique features of the nonprofit environment. The program offers many kinds of support to its members, is inclusive of all kinds of “scholarship,” and aims to be accessible to those who lack time, resources, or formal research training. This culture of scholarship means that Scholars have a support network when they need it, empowering them in more ways than a single blog post can ever capture.
This research was made possible with funding and support from Mitacs, The United Way, SFU Library, and the ScholCommLab. To find out more about the Community Scholars Program Evaluation, you can read the full report here and access the CSP librarians' response here.
ScholCommLab co-director receives funding to analyze patterns of how research data is cited and reused
Stefanie Haustein wins Alfred P. Sloan Foundation research grant of $200K US
Have you ever wondered what motivates researchers to reuse open data and what makes them cite (or not cite) datasets in their work? Or how sharing, reusing and citing open data differs between research areas or changes during a researcher’s career?
ScholCommLab co-director Stefanie was awarded $199,929 US ($281,660 CDN) by the Alfred P. Sloan Foundation to fund Meaningful Data Counts, an interdisciplinary project exploring a range of questions about scholarly data use and citation. Together with co-PI Isabella Peters, Professor of Web Science at the ZBW Leibniz Information Centre for Economics and CAU Kiel University (Germany), Stefanie and her team will conduct the basic research necessary to understand how datasets are viewed, used, cited, and remixed.
“Many researchers have to publish a paper to receive credit for collecting, cleaning, and curating scientific data, because datasets don’t count when applying for a tenure-track job or promotion.”
“Many researchers have to publish a paper to receive credit for collecting, cleaning, and curating scientific data, because datasets don’t count when applying for a tenure-track job or promotion,” Stefanie explains. While an increasing number of journals and funding agencies now require data to be open, research datasets are still considered second-class research contributions. By documenting the uptake and reuse of data across disciplines, she hopes the project will help overcome this perception and incentivize open research practices. “Data metrics that capture dataset views, downloads and citations might help to showcase the value of open data and contribute to elevate its status to first-class scholarly output,” she says.
On the other hand, Stefanie acknowledges that metrics could harm the scientific system in the long run, replicating some of the adverse effects associated with current bibliometric indicators. Therefore, a primary goal of the project is to support the creation of meaningful multidimensional metrics instead of promoting raw numbers and crude rankings. “We want to support the evidence-based development of metrics that are appropriate and well thought-out instead of repeating mistakes of the past by creating more flawed indicators, like the h-index and impact factor,” she explains.
The project team—which, in addition to Stefanie and Isabella, includes collaborators Daniella Lowenberg, Felicity Tayler, Rodrigo Costas, Peter Kraker, Philippe Mongeon, and Nicolas Robinson-Garcia—will therefore take a mixed-methods approach to explore how datasets are (re)used and cited across disciplines. Using metadata provided by DataCite, it will build on ScholCommLab member Asura Enkhbayar’s doctoral research to develop a theory of data citations. As part of the larger Make Data Count (MDC) team, the researchers will work in close collaboration with MDC's infrastructure and outreach specialists. Also funded by Sloan, this project aims to build a bibliometrics dashboard and drive adoption of standardized practices.
“We are thrilled to expand our Make Data Count team and scope to include the bibliometrics community,” says Daniella Lowenberg, the project’s PI and Data Publishing Product Manager at the California Digital Library. Martin Fenner, Technical Director at DataCite and MDC technical lead, agrees: “Going forward, Stefanie and her colleagues will provide us with important insights into researcher behavior towards open data,” he says. “I am excited to see Make Data Count transition into the next phase.”
During the two-year funding period the grant will support two postdoctoral researchers—one at the University of Ottawa (quantitative) and one at Isabella’s Web Science research group at the ZBW. “I am very happy to join forces with the ScholCommLab and strengthen the German-Canadian research connection,” she says. “After having worked together with Stefanie for more than a decade—we studied together in Düsseldorf—this is our first shared research grant, which is really exciting.”
Stefanie agrees: "There's a lot of research to be done, but I'm hopeful that, together with MDC, this project will make a real difference in how the scholarly community values open data."
Interested in learning more? For updates and job postings about the Meaningful Data Counts project, sign up for the ScholCommLab’s newsletter.